Journal of Cancer Research & Therapy
An International Peer-Reviewed Open Access Journal
ISSN 2052-4994


- Download PDF
- |
- Download Citation
- |
- Email a Colleague
- |
- Share:
-
- Tweet
-

Journal of Cancer Research & Therapy
Volume 1, Issue 3, May 2013, Pages 111-118
Original researchOpen Access
Biomarker screening of oral cancer cell lines revealed sub-populations of CD133-, CD44-, CD24- and ALDH1- positive cancer stem cells
- 1 Department of Biomedical Sciences University of Nevada, Las Vegas - School of Dental Medicine, Las Vegas, Nevada, 89106, USA
- 2 Arizona State University, School of Life Sciences, Tempe, Arizona, 85287, USA
- 3 School of Life Sciences University of Nevada, Las Vegas, Las Vegas, Nevada, 89154, USA
*Corresponding author: Kingsley K, Department of Biomedical Sciences, University of Nevada, Las Vegas-School of Dental Medicine, 1001 Shadow Lane, Las Vegas, Nevada, 89106, USA, Tel.: 702-774-2623. E-mail: Karl.Kingsley@unlv.edu
Received 10 November 2012 Revised 1 April 2013 Accepted 7 April 2013 Published 14 April 2013
DOI: http://dx.doi.org/10.14312/2052-4994.2013-17
Copyright: © 2013 Kendall K, et al. Published by NobleResearch Publishers. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
AbstractTop
Head and neck squamous cell carcinoma (HNSCC) ranks sixth worldwide for cancer-related mortality. For the past several decades the mainstay of treatment for HNSCC has been surgery and external beam radiation, although more recent trials combining chemotherapy and radiation have demonstrated improvements. However, cancer recurrence and treatment failures continue to occur in a significant percentage of patients. Recent advances in tumor biology have led to the discovery that many cancers, including HNSCC, may contain subpopulations of cells with stem cell-like properties that may explain relapse and recurrence. The objective of this study was to screen existing oral cancer cell lines for biomarkers specific for cells with stem cell-like properties. RNA was isolated for RT-PCR screening using primers for specific mRNA of the biomarkers: CD44, CD24, CD133, NANOG, Nestin, ALDH1, and ABCG2 in CAL27, SCC25 and SCC15 cells. This analysis revealed that some oral cancer cell lines (CAL27 and SCC25) may contain small subpopulations of adhesion- and contact-independent cells (AiDC) that also express tumor stem cell markers, including CD44, CD133, and CD24. In addition, CAL27 cells also expressed the intracellular tumor stem cell markers, ALDH1 and ABCG2. Isolation and culture of the adhesion- and contact-independent cells from CAL27 and SCC25 populations revealed differential proliferation rates and more robust inhibition by the MEK inhibitor PD98059, as well as the chemotherapeutic agents Cisplatin and Paclitaxel, within the AiDC CAL27 cells. At least one oral cancer cell line (CAL27) contained subpopulations of cells that express specific biomarkers associated with tumor stem cells which were morphologically and phenotypically distinct from other cells within this cell line.
Keywords: oral cancer; stem cell; biomarker
IntroductionTop
Head and neck squamous cell carcinoma (HNSCC) is among the most common cancers worldwide and currently ranks sixth for cancer-related mortality [1, 2]. For the past several decades the mainstay of treatment for HNSCC has been surgery and external beam radiation [1], although recent clinical trials have demonstrated benefit from combining chemotherapy and radiation for advanced stage disease [3-8]. However, HNSCC recurrence and treatment failures continue to occur in a significant percentage of patients. The biological mechanisms that underlie why some HNSCCs respond favorably to treatment and others do not are under active investigation, but much remains to be elucidated [9-13].
Some evidence has suggested many therapies fail to prevent cancer relapse and metastasis because of small, surviving populations of cancer stem cells (CSC) [14, 15]. Studies in a wide variety of malignancies, including those in the breast, brain, lung, liver, pancreas and colon, have demonstrated that distinct sub-populations of CSC maintain the ability to undergo self-renewal and differentiation and hence have the capacity not only to initiate tumorigenesis and support ongoing tumor growth, but also to survive many traditional therapeutic interventions [16-19]. As many tumor types appear to contain heterogeneous populations of cancer cells, the ever-growing corpus of research now suggests that CSCs may be responsible not only for tumorigenesis, but also may be a significant contributing factor in resistance to cancer therapies [20-22].
Advances in tumor biology have also led to the recent discovery that HNSCCs may also be supported by cells with stem-like properties, although this research is far from conclusive [23-25]. Most studies have focused on the identification of these sub-populations or side populations (SP) of CSC in primary explants from HNSCCs [26-28]. Many of these studies used previously validated cell surface CSC biomarkers, including CD44 expression, for isolation and characterization – although these HNSCC biomarkers have expanded to include CD133 and CD24 [29-31]. CD44 is a cell surface glycoprotein with well-characterized functions, which include cell-cell interactions, participation in cellular adhesion and migration, and as a receptor for hyaluronic acid [32-34]. CD44 triggers receptor tyrosine kinases in many cancers, such as the cell-cycle promoting Src/STAT3, and c-myc/MAPK pathways, while new evidence has suggested the C-terminus may play an active role in tumor sphere formation among CSC [35]. CD133 or prominin-1 is a 5-transmembrane glycoprotein that appears to localize to cellular protrusions and is differentially expressed not only in hematopoietic stem cells, but also in CSC isolated from solid tumors [36, 37]. Moreover, the cytoplasmic portion of CD133 has recently been identified as a novel binding partner of Src, and is also phosphorylated by Src kinase as are other cytosolic targets (Ras/MAPK, c-myc/MAPK), which suggests that both CD44 and CD133 activate, transduce, and amplify common downstream cell-cycle signaling pathways [38-40]. CD24 is a cell surface sialoglycoprotein and cancer stem cell marker that triggers expression of Oct4, Sox2, NANOG, and Nestin transcription factors in CSC, similar to CD44 and CD133 [41-45].
In addition to reports of intracellular biomarkers for CSC up-regulated by CD133, and CD24, such as Oct4, Sox, NANOG , and Nestin, newer reports have demonstrated that aldehyde dehydrogenase (ALDH) in combination with CD44 may serve to identify and define CSC [42, 46-48]. ALDH1 is part of a larger family of NADP-dependent enzymes that function to assist tumor resistance to chemotherapy and correlates with HNSCC CSC expression of Oct4, Sox2, and Nanog [48, 49]. In fact, this evidence has suggested that co-expression of CD44 and ALDH is relatively rare (1-3% of tumor cells) and may identify those cells uniquely capable of generating tumors. New evidence now suggests that restricted expression of CD44 and ALDH1 co-localizes with ATP-binding cassette (ABC) transporter gene expression [46]. ABC transporters are expressed in chemoresistant stem cell phenotype-expressing tumor initiating cells that function as drug efflux transporters for doxorubicin and 5-FU [50, 51] and are closely related to ABCB1, ABCC5, ABCC6, and ABCG2 that function as efflux pumps for Cisplatin and are predictive for OSCC chemosensitivity [52-55].
A growing number of studies have identified and isolated CSC from HNSCC and OSCC. However, heterogeneous responsiveness in SPs of CSCs has been found among these primary tumor explants [56, 57]. These commonly observed variations within primary tissues strongly suggest the need for more extensive research using well-characterized HNSCC cell lines to identify novel biomarkers and their correlations with treatment responsiveness [23, 29]. The purpose of this study was to screen previously untested oral cancer cell lines to assess them for heterogeneous populations that may suggest the presence of CSC SPs within these well-characterized HNSCC lines [23, 58, 59].
Materials and methodsTop
Cell lines
The human oral scquamous cell carcinoma (OSCC) cell lines used in this study, CAL27
(CRL-2095), SCC15 (CRL-1623), and SCC25 (CRL-1628) were obtained from American Type
Culture Collection (ATCC: Manassas, VA). CAL27 cells were maintained in Dulbecco’s
Modified Eagle’s Medium (DMEM) with 4 mM L-glutamine, adjusted to contain 3.7 g/L
sodium bicarbonate and 4.5 g/L glucose from Hyclone (Logan, UT). SCC15 and SCC25
cells were maintained in a 1:1 mixture of DMEM and Ham’s F12 medium with 2.5 mM
L-glutamine, modified to contain 15 mM HEPES, 0.5 mM sodium pyruvate, and 1.2 g/L
sodium bicarbonate (ATCC), supplemented with 400 ng/ml hydrocortisone from Sigma-Aldrich
(St. Louis, MO). Media for all cell lines were supplemented with 10% fetal bovine
serum (FBS), and with 1% Penicillin (10,000 units/mL)-Streptomycin (10,000 µg/mL)
solution (HyClone). Cell cultures were maintained in 75 cm2 Becton, Dickinson
(BD) Falcon tissue-culture treated flasks (Bedford, MA) at 37 °C and 5% CO2
in humidified chambers.
Passage number
All dividing cells undergo spontaneous mutation at a near constant rate of approximately
10-9 per nucleotide, therefore extended in vitro cell culture and clonal
expansion can lead to the emergence of new genotypes and altered cellular phenotypes
over time [60-62]. The main alterations observed in cultured human cancers include
alterations in chromosomal copy number, mitochondrial DNA sequence changes, and
vast alterations in gene promoter methylation (and therefore, gene expression).
Based upon this evidence, all experiments were performed on early passage cells
(between passage three and ten (P3-P10); CAL27 between passages four and passage
eight (P4-P8); SCC15 between passages three and six (P3-P6); SCC25 between passages
five and passage ten (P4-P10).
Microscopy and staining
Cells were visualized with a Zeiss Axiovert 40 inverted microscope (Gottingen, Germany),
and images were captured at 200X magnification with a Canon PowerShot G6 digital
camera (Tokyo, Japan). Digital images were subsequently processed using Adobe Photoshop
(San Jose, CA) Image Analysis tools. In brief, some cell cultures were fixed using
50 µL of 10% buffered formalin and subsequently stained using crystal violet
1% aqueous solution to document cell morphology, cell spreading and confluence.
RNA isolation
RNA was isolated from 1.5 x 107 cells of each of the experimental cell
lines, using AB gene Total RNA Isolation Reagent (Epsom, Surrey, UK) in accordance
with the procedure recommended by the manufacturer for RT-PCR analysis. RNA concentration
and purity were calculated using UV spectroscopy. The absorbance of diluted RNA
samples (10 µL of RNA sample in 490 µL nuclease-free water, pH 7.0)
was measured at 260 and 280 nm. RNA purity was determined by calculating the ratio
of A260:A280, which should be >1.80. Concentration for RNA samples was determined
by the A260 reading of 1 =40 µg/mL RNA, based on an extinction coefficient
calculated for RNA in nuclease-free water. Concentration was calculated as 40 x
A260 absorbance measure x dilution factor [50]. Total yield was determined by concentration
x sample volume in mL.
Example: RNA standard
A260 =0.75
Concentration = 40 x 0.75 x 50 = 1,500 µg/mL
Yield = 1,500 µg/mL x 1.0 mL = 1,500 µg or 1.5 mg RNA
Similar RNA concentrations were obtained from each cell line, which ranged from
876 – 955 ng/µL. Analysis of A260/A280 ratio confirmed the purity, which ranged
between 1.66 and 1.88.
Reverse-transcription polymerase chain reaction (RT-PCR)
To quantify the expression of CSC-specific mRNA, RT-PCR was performed on total RNA
using the ABgene Reverse-iT One-Step RT-PCR Kit (ReadyMix Version) and a Mastercycler
gradient thermocycler (Eppendorf: Hamburg, Germany) using the following primers
synthesized by SeqWright (Houston, TX):
CD44 FORWARD: GAAAGGCATCTTATGGATGTGC;
CD44 REVERSE:
CTGTAGTGAAACACAACACC;
CD133 FORWARD: CTCATGCTTGAGAGATCAGGC;
CD133 REVERSE: CGTTGAGGAAGATGTGCACC;
CD24 FORWARD: ACTCTCACTTGAAATTGGGC;
CD24 REVERSE: GCACATGTTAATTACTAGTAAAGG;
c-myc FORWARD: TCCAGCTTGTACCTGCAGGATCTGA;
c-myc REVERSE: CCTCCAGCAGAAGGTGATCCAGACT;
STAT3 FORWARD: AGACTACAGGCCCTCAGCAA;
STAT3 REVERSE: CCTCTGTCAGGAAAGGCTTG;
Nestin FORWARD: CGTTGGAACAGAGGTTGGAG;
Nestin REVERSE: TCCTGAAAGCTGAGGGAAG;
NANOG FORWARD: GCTGAGATGCCTCACACGGAG;
NANOG REVERSE: TCTGTTTCTTGACTGGGACCTTGTC;
Oct4 FORWARD: TGGAGAAGGAGAAGCTGGAGCAAAA;
Oct4 REVERSE: GGCAGATGGTCGTTTGGCTGAATA;
Sox2 FORWARD: ATGGGCTCTGTGGTCAAGTC;
Sox2 REVERSE:
CCCTCCCAATTCCCTTGTAT;
ALDH1 FORWARD: GACACGGATCCATGTTGCGCGCTGCCGCCCGCTTCGG;
ALDH1 REVERSE: GACACGAATTCTTATGAGTTCTTCTGAGGCACTTTGAC;
ABCG-2 FORWARD: AGTTCCATGGCACTGGCCATA;
ABCG-2 REVERSE: TCAGGTAGGCAATTGTGAAGG;
GAPDH FORWARD: ATCTTCCAGGAGCGAGATCC;
GAPDH REVERSE: ACCACTGACACGTTGGCAGT;
RNA standard: GAPDH
RNA standards obtained from standardized control cells, human gingival fibroblasts
(0.3-0.5 x 106 cells/mL were used to establish the minimum threshold
(CT) and saturation (CS) cycles required for calibration and concentration comparisons
using relative endpoint (RE) RT-PCR. GAPDH signal detection above background or
CT required a minimum of ten cycles (C10), with saturation or CS observed at C40.
Based upon these data, RE-PCR was performed at C35, above the lower detection limit
but below the saturation limit.
One µg of template (total) RNA was used for each reaction. The reverse transcription step ran for 30 min. at 47°C, followed by denaturation for 2 min. at 94°C. Thirty-five amplification cycles were run, consisting of 20 sec. denaturation at 94°C, 30 sec. of annealing at 58°C, and 6.5 min. of extension at 72°C. Final extension was run for 5 min. at 72°C. Reaction products were separated by gel electrophoresis using Reliant 4% NuSieve® 3:1 Plus Agarose gels (Lonza: Rockland, ME). Bands were visualized by UV illumination of ethidium-bromide-stained gels and captured using a Kodak Gel Logic 100 Imaging System and 1D Image Analysis Software (Eastman Kodak: Rochester, NY). Quantitation of RT-PCR band densitometry and relative mRNA expression levels were performed using Adobe Photoshop (San Jose, CA) imaging software, Image Analysis tools.
Proliferation
Proliferation assays were performed in the appropriate complete media in Corning
Costar 96-well assay plates (Corning, NY) at a concentration of 1.2 x 104
cells per well, and proliferation was measured over three days. Cultured cells were
fixed at three time points, after 24 hrs (day 1), after 48 hrs (day 2), and after
72 hrs (day 3) using 50 µL of 10% buffered formalin, and were stained with
crystal violet 1% aqueous solution (Fisher Scientific: Fair Lawn, NJ). The relative
absorbance was measured at 630 nm using a Bio-Tek ELx808 microplate reader (Winooski,
VT). Data were analyzed and graphed using Microsoft Excel (Redmond, WA) and SPSS
(Chicago, IL). Three separate, independent replications of each experiment were
performed.
PD98059, Cisplatin, and Paclitaxel effects on adhesion-independent cell tumor spheres
(AiDC-TS)
A simple method to effectively separate morphologically distinct tumor spheres employs
the use of a cell scraper to dislodge the desired cells for subsequent transfer
using a micropipette into a new culture plate for experimentation [29]. Proliferation
assays were setup as described above with one addition; where applicable, the cell
culture medium was supplemented with PD98059 (50 µM), a MEK1 inhibitor from
Caliochem/EMD Biosciences/Millipore, M.W. 267.3 (Darmstadt, Germany); with Paclitaxel
(1.25 µg/mL), a mitotic inhibitor from MP Biomedicals, MW 853.9 (Santa Ana,
CA) often used in cancer chemotherapy and with oral cancers (23,31), or with Cisplatin
(2.0 µg/mL), cis-diamminedichloroplatinum II, a platinum-containing cancer
drug used with oral cancers (63,64) from Selleck Chemical, LLC , M.W. 300.5 (Houston,
TX).
ResultsTop
Cell culture
Oral cancer (CAL27, SCC25 and SCC15) and normal non-tumorigenic (HGF-1, Hs27) cell
lines were maintained and cultured in vitro (Figure 1). This analysis revealed
that two oral cancer cell lines, CAL27 (Figure 1A) and SCC25 (Figure 1B) contained
a small number of cell sub-populations that were phenotypically distinct from the
surrounding cells. These cells formed adhesion-independent and cell-cell contact-independent
tumor spheres that continued to grow despite confluence of the surrounding monolayer.
There was no evidence of these structures or subpopulations in any other cell line
examined, which included the oral cancer SCC15, (Figure 1C) and both normal oral
gingiva HGF-1, (Figure 1D) and fibroblast Hs27 (Figure 1E) cell lines.
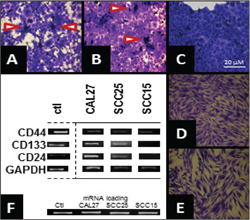
Figure 1 Morphology and cell surface markers of oral cancer cell lines reveal stem cell-like subpopulations. In vitro cell culture of oral cancer (A-C) and normal (D-E) cell lines revealed CAL27 (A) and SCC25 (B) cell lines harbor smaller subpopulations that appear to grow in an adhesion- and cell contact-independent manner, forming small tumor spheres above the monolayer; this was not observed in SCC15 (C) or normal oral gingiva HGF-1 (D) or fibroblast Hs27 (D) controls. (F) RT-PCR analysis on total RNA isolated from the oral cell lines revealed expression of mRNA for tumor markers (CD44) and expression of the tumor stem cell markers CD133 and CD24 in CAL27 cells, and to a lesser extent in SCC25 cells (but not SCC15).
RT-PCR
To analyze whether these cell lines (or subpopulations thereof) expressed biomarkers
indicative of cancer stem cells, RNA was isolated from 1.5 x 107 cells
from each cell line for RT-PCR analysis (Figure 1F). This analysis revealed all
three oral cancer cell lines expressed mRNA for the non-specific tumor marker CD44.
Expression for tumor stem cell surface markers CD133 and CD24 was noted in CAL27,
and to a lesser extent in SCC25, but not SCC15, cells.
Intracellular markers
Total RNA from each cell line was used to assess if there was differential expression
of specific, intracellular tumor stem cell markers in the cell contact-, adhesion-independent
subpopulations observed in some oral cancer cell lines (Figure 2). All cell lines
expressed mRNA specific for the cell cycle promoter c-myc, as well as the oral cancer
growth promoter STAT3. However, the expression of the tumor stem cell markers Nestin,
NANOG, Sox2, and Oct4 were observed primarily in CAL27 and SCC25 cell lines and
were either weakly or not expressed in the SCC15 cell line. Interestingly, expression
of ALDH1, as well as the ABC transporter ABCG2 was only observed within the CAL27
cell line.
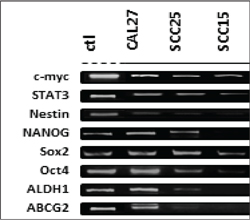
Figure 2 mRNA analysis of intracellular markers using total RNA. RT-PCR was performed on total RNA extracted from CAL27, SCC25 and SCC15 cells. Relative endpoint (RE) RT-PCR revealed that each cell line expressed mRNA for the tumor suppressor p53, as well as the oral cancer promoters, c-myc, ODC and STAT3. The marker for lower survival p27, as well as the tumor stem cell marker Nestin, were observed only in CAL27 cells.
Morphology and growth inhibition
Subpopulations of CAL27 cells were isolated to determine if phenotypic and morphologic
distinctions could be assessed in vitro. Separation of potential tumor spheres
was facilitated with a cell scaper for transfer into fresh plates for morphological
analysis (Figure 3). The resulting separation revealed the remaining CAL27 adhesion-dependent
cells (ADC) growing in uniform monolayers (Figure 3A), while the isolated adhesion-independent
tumor spheres (AiDC-TS) grew in cell clusters that were not contact or adhesion-dependent
within the monolayer (Figure 3B). Three day growth assays revealed the CAL27-AiDC-TS
cells (Figure 3D) grew more rapdily than the CAL27-ADC (Figure 3C) at all time points
measured. In addition, administration of PD98059, a potent and selective inhibitor
of MAP kinase kinase (MEK kinase) inhibited both CAL27-ADC and CAL27-AiDC-TS growth,
although this inhibition effect was more pronounced in CAL27-AiDC-TS. The addition
of Paclitaxel was sufficient to inhibit growth of CAL27-ADC, although this was not
statistically different from the inhibition observed under PD98059 administration
(-18.4%, -22.4%, respectively). Cisplatin, however, strongly inhibited CAL27-ADC
growth (-38.8%), which was more robust than the inhibition observed with PD98059
or Paclitaxel administration (p<0.05). It was also observed that CAL27-AiDC-TS,
which were strongly inhibited by PD98059 administration (-34.9%), were also inhibited
by Paclitaxel administration (-9.3%) – although this inhibition was comparably lower
and was comparably less robust than with the CAL27-ADC (p<0.05). Moreover, the
administration of Cisplatin also inhibited CAL27-AiDC-TS cel growth (-19.8%), which
was more robust than observed with Paclitaxel, but less than the inhibition induced
by PD98059.

Figure 3 CAL27 subpopulations reveal distinct morphologies and growth inhibition. Separation of adhesion dependent cells (ADC) and adhesion-independent cell tumor spheres (AiDC-TS) from CAL27 cell cultures revealed smooth adhesion and contact-dependent cells (A) and adhesion-independent, contact-independent tumor spheres. (B) 96-well growth assays of these isolates revealed CAL27-AiDC-TS proliferation was 53.6% greater than CAL27-ADC. (C) Inhibition of growth using Cisplatin was more robust than Paclitaxel or the MEK inhibitor PD98059 among CAL27-ADC cells. (D) Although observed growth was more rapid, in general, growth inhibition among CAL27-AiDC-TS was greatest under PD98059 administration, than either Paclitaxel or Cisplatin.
To facilitate a quantitative comparison of growth inhibition induced by PD98059, Paclitaxel, and Cisplatin, the percentage of growth inhibition among CAL27-ADC and CAL27-AiDC-TS was graphed (Figure 4A). These results outline the much stronger inhibiting effects of Cisplatin on CAL27-ADC compared with either Paclitaxel or PD98059. Although both chemotherapeutic agents (Paclitaxel, Cisplatin) inhibited growth among CAL27-AiDC-TS, this was lower than the inhibition observed with these same agents on CAL27-ADC. Finally, PD98050 also inhibited CAL27-AiDC-TS growth, although this inhibition was much more pronounced than with CAL27-ADC and much more robust than the inhibition by Paclitaxel or Cisplatin.
To assess any significant differences in mRNA expression, RNA was isolated from CAL27-ADC and CAL27-AiDC-TS for comparison (Figure 4B). No observable differences were noted in mRNA expression of cell-cycle regulators c-myc or STAT3. Some minor differences were observed among the CSC biomarkers, Nestin and NANGO, although more striking differences were observed in the mRNA expression of the drug efflux transporter ABCG2, as well as ALDH1, and both Sox2 and Oct4.

Figure 4 CAL27-ADC and CAL27-AiDC-TS exhibit differential responses. (A) Comparison with baseline growth (without inhibitors) revealed no significant differences in growth inhibition among CAL27-ADC under PD98059 (-22.4%) or Paclitaxel (-28.4%), but much stronger inhibition with Cisplatin (-38.8%). CAL27-AiDC-TS were also inhibited by Paclitaxel (-9.3%) and Cisplatin (-19.8%), although this was more pronounced with PD98059 (-34.9%).
DiscussionTop
This study found that of three previously unscreened oral squamous cell carcinoma cell lines (CAL27, SCC25, SCC15), two may harbor stem-cell like subpopulations. These two cell lines (CAL27, SCC25) express mRNA typically associated with cancer stem cells, including CD133, CD44 and CD24, as well as intracellular cancer stem cell biomarkers such as NANOG and Nestin. In addition, expression of ALDH1 and ABCG2 were observed only in CAL27 cells, with much higher expression among the CAL27-AiDC-TS isolates, suggesting that this cell line may be the most likely to harbor a subpopulation of CSC.
These findings complement the growing body of evidence suggesting that oral cancer cell lines may, in fact, harbor subpopulations of cancer stem cells [23, 29, 30, 50, 59]. Recent evidence suggests that these subpopulations may persist or even further develop in well-established, commercial cell lines and cancer tissue explants, such as SCC-4, SCC-9, PCI-4A, PCI-8, PCI-9A, PCI-13 – part of the newest expanding field of oral tumor biology research [23, 28, 31]. Perhaps the screening of all commercially available cell lines for these CSC subpopulations may provide more information about their existence and more opportunities to analyze these phenotypes and their response behavior to anti-tumor therapies in future research.
The finding that adhesion-independent tumor sphere forming cell colonies from the CAL27 cell line display a differential response to the MEK inhibitor PD98059, as well as to the chemotherapeutic agents Cisplatin and Paclitaxel, may provide new evidence of a potential mechanism of previously observed phenomenon. For example, many of the traditional HNSCC therapies are designed specifically to target rapidly dividing cancer cells [3-8]. However, many studies now demonstrate that cancer stem cells may respond differentially to traditional chemotherapies by down-regulating proliferation, which suggest new highly-selective therapies to target cancer stem cells are needed in conjunction with traditional chemo- and radiation therapy [59, 66, 67] to target these cancer cell sub-populations. That CAL27 adhesion independent cell growth is inhibited more significantly by PD98059, than by Cisplatin or Paclitaxel, may suggest that growth inhibition response among this subpopulation of cells may be one of many mechanisms that allow for the increased survival and treatment resistance observed during clinical chemotherapy. Furthermore, that CAL27-AiDC-TS are less responsive to Cisplatin than CAL27-ADC may suggest the need for experimental studies on the effects of co-administration of traditional chemotherapy with complementary cancer stem cell therapies or the development of new treatment protocols that target both primary tumor cells and stem cells.
There are limitations involved in this type of in vitro work, similar to many other types of pre-clinical research, that utilize oral cancer cell lines. One of the most significant is the possibility that each cell line may harbor genetic mutations or other genetic anomalies that could alter the experimental outcomes. For example, previous research has revealed that the SCC25 cell line harbors a deletion in a critical cell cycle regulator, cyclin dependent kinase, Cdk1 [68]. In addition, the SCC15 cell line is known to harbor a single nucleotide polymorphism (SNP) that has been demonstrated to influence the expression and efficacy of the tumor suppressor, S100A2 – another potential confounder of proliferation of growth studies [69]. Finally, CAL27 cells are known to harbor a nonsense mutation in SMAD4, which modulates the transforming growth factor (TGF) signaling pathway and cell growth [70]. More recent evidence now suggests that, in addition to genetic mutations and deletions, many cells may dysregulate expression of many genes due to dypermethylation events rather than due to fixed genetic alterations [71].
ConclusionTop
Despite these limitations, these results provide clear evidence that oral cancer cell lines may harbor subpopulations of cancer stem cells, which may be sufficient to alter the response of each cell line to chemotherapy or other treatments. In addition, the isolation and characterization of these cancer stem cells allows for the possibility that novel treatment regimens that incorporate standard chemotherapy in combination with specific cancer cell inhibitors can be assessed for concentration and dose to determine clinical relevance. This study is among the first to demonstrate these effects in well characterized cell lines and provide a model for further in-depth evaluation of the therapeutic possibility of new combination treatments for patients with oral cancer.
Funding
This research was supported, in part, by an American Cancer Society Institutional
Research Grant to KK (ACS#103719).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial
or financial relationships that could be construed as a potential conflict of interest.
Acknowledgements
The authors would like to acknowledge the UNLV-SDM Office of Research for their
support.
ReferencesTop
[1] Haddad RI, Shin DM (2008) Recent advances in head and neck cancer. N Engl J Med 359:1143–1154. Article Pubmed
[2] Rothenberg SM, Ellisen LW (2012) The molecular pathogenesis of head and neck squamous cell carcinoma. J Clin Invest 122:1951–1957. Article Pubmed
[3] Bae WK, Hwang JE, Shim HJ, Cho SH, Lee KH, et al. (2013). Multicenter phase II study of weekly docetaxel, cisplatin, and S-1 (TPS) induction chemotherapy for locally advanced squamous cell cancer of the head and neck. BMC Cancer 13:102. Article Pubmed
[4] Hsu HW, Gridley DS, Kim PD, Hu S, de Necochea-Campion R, et al. (2013) Mirshahidi S. Linifanib (ABT-869) enhances radiosensitivity of head and neck squamous cell carcinoma cells. Oral Oncol S1368–8375(13)00041–00049. Article Pubmed
[5] Martins RG, Parvathaneni U, Bauman JE, Sharma AK, Raez LE, et al. (2013) Cisplatin and Radiotherapy With or Without Erlotinib in Locally Advanced Squamous Cell Carcinoma of the Head and Neck: A Randomized Phase II Trial. J Clin Oncol 31:1415–1421. Article Pubmed
[6] Epstein JB, Zhang L, Rosin M (2002) Advances in the diagnosis of oral premalignant and malignant lesions. J Can Dent Assoc 68: 617–621. Article Pubmed
[7] Farah CS, McCullough MJ (2008) Oral cancer awareness for the general practitioner: new approaches to patient care. Aust Dent J 53: 2–10. Article Pubmed
[8] da Silva SD, Hier M, Mlynarek A, Kowalski LP, Alaoui-Jamali MA (2012) Recurrent oral cancer: current and emerging therapeutic approaches. Front Pharmacol 3:149. Article Pubmed
[9] Epstein JB, Thariat J, Bensadoun RJ, Barasch A, Murphy BA, et al. (2012) Oral complications of cancer and cancer therapy: from cancer treatment to survivorship. CA Cancer J Clin 62:400–422. Article Pubmed
[10] Siegel R, Naishadham D, JemalA (2012) Cancer statistics, 2012. CA Cancer J Clin 62:10–29. Article Pubmed
[11] Bauman JE, Michel LS, Chung CH (2012) New promising molecular targets in head and neck squamous cell carcinoma.CurrOpinOncol 24:235–242. Article Pubmed
[12] Ho AS, Kraus DH, Ganly I, Lee NY, Shah JP, et al. (2013) Decision making in the management of recurrent head and neck cancer. Head Neck doi: 10.1002/hed.23227. Article Pubmed
[13] Hunt JL, Barnes L, Lewis JS Jr, Mahfouz ME, Slootweg PJ, et al. (2013) Molecular diagnostic alterations in squamous cell carcinoma of the head and neck and potential diagnostic applications. Eur Arch Otorhinolaryngol. Article Pubmed
[14] Wicha MS, Liu S, Dontu G (2006) Cancer stem cells: an old idea--a paradigm shift. Cancer Res 66:1883–1890. Article Pubmed
[15] Tirino V, Desiderio V, d'Aquino R, De Francesco F, Pirozzi G, et al. (2008) Detection and characterization of CD133 cancer stem cells in human solid tumours. PLoS One 3:e3469 doi: 10.1371/journal.pone.0003469. Article Pubmed
[16] Singh SK, Clarke ID, Terasaki M, Bonn VE, Hawkins, et al. (2003) Identification of a cancer stem cell in human brain tumors. Cancer Res 63:5821–5828. Article Pubmed
[17] Dontu G, El-Ashry D, Wicha MS (2004) Breast cancer, stem/progenitor cells and the estrogen receptor.Trends Endocrinol Metab 15:193–197. Article Pubmed
[18] Kim CF, Jackson EL, Woolfenden AE, Lawrence S, Babar I, et al. (2005) Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell 121:823–835. Article Pubmed
[19] Li C, Heidt DG, Dalerba P, Burant CF, Zhang L, et al. (2007) Identification of pancreatic cancer stem cells. Cancer Res 67:1030–1037. Article Pubmed
[20] Rosen JM, Jordan CT (2009) The increasing complexity of the cancer stem cell paradigm. Science 324: 1670–1673. Article Pubmed
[21] Bohl SR, Pircher A, Hilbe W (2011) Cancer stem cells: characteristics and their potential role for new therapeutic strategies. Onkologie 34:269–274. Article Pubmed
[22] Clevers H (2011) The cancer stem cell: premises, promises and challenges. Nat Med 17:313–319. Article Pubmed
[23] Felthaus O, Ettl T, Gosau M, Driemel O, Brockhoff G (2011) Cancer stem cell-like cells from a single cell of oral squamous carcinoma cell lines.BiochemBiophys Res Commun 407: 28–33. Article Pubmed
[24] Yanamoto S, Kawasaki G, Yamada S, Yoshitomi I, Kawano T (2011) Isolation and characterization of cancer stem-like side population cells in human oral cancer cells. Oral Oncol 47:855–860. Article Pubmed
[25] Albers AE, Chen C, Köberle B, Qian X, Klussmann JP, et al. (2012) Stem cells in squamous head and neck cancer. Crit Rev Oncol Hematol 8: 224–240. Article Pubmed
[26] Prince ME, Sivanandan R, Kaczorowski A, Wolf GT, Kaplan MJ, et al. (2007) Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc Natl Acad Sci U S A 104:973–978. Article Pubmed
[27] Okamoto A, Chikamatsu K, Sakakura K, Hatsushika K, Takahashi G, et al. (2009) Expansion and characterization of cancer stem-like cells in squamous cell carcinoma of the head and neck. Oral Oncol 45:633–639. Article Pubmed
[28] Zhang P, Zhang Y, Mao L, Zhang Z, Chen W (2009) Side population in oral squamous cell carcinoma possesses tumor stem cell phenotypes. Cancer Lett 277:227–234. Article Pubmed
[29] Harper LJ, Piper K, Common J, Fortune F, Mackenzie IC (2007) Stem cell patterns in cell lines derived from head and neck squamous cell carcinoma. J Oral Pathol Med 36:594–603. Article Pubmed
[30] Chiou SH, Yu CC, Huang CY, Lin SC, Liu CJ, et al. (2008) Positive correlations of Oct-4 and Nanog in oral cancer stem-like cells and high-grade oral squamous cell carcinoma. Clin Cancer Res 14:4085–4095. Article Pubmed
[31] Zhang Q, Shi S, Yen Y, Brown J, Ta JQ, et al. (2010) A subpopulation of CD133() cancer stem-like cells characterized in human oral squamous cell carcinoma confer resistance to chemotherapy. Cancer Lett 289:151–160. Article Pubmed
[32] Misra S, Hascall VC, Berger FG, Markwald RR, Ghatak S (2008) Hyaluronan, CD44, and cyclooxygenase-2 in colon cancer. Connect Tissue Res 49:219–224. Article Pubmed
[33] Ghatak S, Misra S, Toole BP (2005) Hyaluronan constitutively regulates ErbB2 phosphorylation and signaling complex formation in carcinoma cells. J Biol Chem 280:8875–8883. Article Pubmed
[34] Misra S, Toole BP, Ghatak S (2006) Hyaluronan constitutively regulates activation of multiple receptor tyrosine kinases in epithelial and carcinoma cells. J Biol Chem 281:34936–34941. Article Pubmed
[35] Su YJ, Lai HM, Chang YW, Chen GY, Lee JL (2011) Direct reprogramming of stem cell properties in colon cancer cells by CD44. EMBO J 30:3186–3199. Article Pubmed
[36] Corbeil D, Röper K, Hellwig A, Tavian M, Miraglia S, et al. (2000) The human AC133 hematopoietic stem cell antigen is also expressed in epithelial cells and targeted to plasma membrane protrusions. J Biol Chem 275:5512–5520. Article Pubmed
[37] O'Brien CA, Pollett A, Gallinger S, Dick JE (2007) A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 445:106–110. Article Pubmed
[38] Chen YS, Wu MJ, Huang CY, Lin SC, Chuang TH, et al. (2011) CD133/Src axis mediates tumor initiating property and epithelial-mesenchymal transition of head and neck cancer. PLoS One 6:e28053 doi: 10.1371/journal.pone.0028053. Article Pubmed
[39] Boivin D, Labbé D, Fontaine N, Lamy S, Beaulieu E, et al. (2009) The stem cell marker CD133 (prominin-1) is phosphorylated on cytoplasmic tyrosine-828 and tyrosine-852 by Src and Fyn tyrosine kinases. Biochemistry 48:3998–4007. Article Pubmed
[40] Guarino M (2010) Src signaling in cancer invasion. J Cell Physiol 223:14–26. Article Pubmed
[41] Castellón EA, Valenzuela R, Lillo J, Castillo V, Contreras HR, et al. (2012) Molecular signature of cancer stem cells isolated from prostate carcinoma and expression of stem markers in different Gleason grades and metastasis. Biol Res 45:297–305. Article Pubmed
[42] Dalley AJ, AbdulMajeed AA, Upton Z, Farah CS (2013) Organotypic culture of normal, dysplastic and squamous cell carcinoma-derived oral cell lines reveals loss of spatial regulation of CD44 and p75 NTR in malignancy. J Oral Pathol Med 42:37–46. Article Pubmed
[43] Kang KS, Choi YP, Gao MQ, Kang S, Kim BG, et al. (2013) CD24⁺ ovary cancer cells exhibit an invasive mesenchymal phenotype. Biochem Biophys Res Commun 432:333–338. Article Pubmed
[44] Zhang C, Li C, He F, Cai Y, Yang H (2011) Identification of CD44+ CD24+ gastric cancer stem cells. J Cancer Res Clin Oncol 137:1679–1686. Article Pubmed
[45] Chen C, Wei Y, Hummel M, Hoffmann TK, Gross M, et al. (2011) Evidence for epithelial-mesenchymal transition in cancer stem cells of head and neck squamous cell carcinoma. PLoS One 6:e16466 doi: 10.1371/journal.pone.0016466. Article Pubmed
[46] Grimm M, Krimmel M, Polligkeit J, Alexander D, Munz A, et al. (2012) ABCB5 expression and cancer stem cell hypothesis in oral squamous cell carcinoma. Eur J Cancer 48:3186–3197. Article Pubmed
[47] Zou B, Sun S, Qi X, Ji P (2012) Aldehyde dehydrogenase activity is a cancer stem cell marker of tongue squamous cell carcinoma. Mol Med Rep 5:1116–1120. Article Pubmed
[48] Chen YC, Chen YW, Hsu HS, Tseng LM, Huang PI, et al. (2009) Aldehyde dehydrogenase 1 is a putative marker for cancer stem cells in head and neck squamous cancer. Biochem Biophys Res Commun 385:307–313. Article Pubmed
[49] Clay MR, Tabor M, Owen JH, Carey TE, Bradford CR, et al. (2010) Single-marker identification of head and neck squamous cell carcinoma cancer stem cells with aldehyde dehydrogenase. Head Neck 32:1195–1201. Article Pubmed
[50] Wilson BJ, Schatton T, Zhan Q, Gasser M, Ma J, et al. (2011) ABCB5 identifies a therapy-refractory tumor cell population in colorectal cancer patients. Cancer Res 71:5307–5316. Article Pubmed
[51] Schatton T, Murphy GF, Frank NY, Yamaura K, Waaga-Gasser AM, et al. (2008) Identification of cells initiating human melanomas. Nature 451:345–349. Article Pubmed
[52] Jovelet C, Bénard J, Forestier F, Farinotti R, Bidart JM, et al. (2012) Inhibition of P-glycoprotein functionality by vandetanib may reverse cancer cell resistance to doxorubicin. Eur J Pharm Sci 46:484–491. Article Pubmed
[53] Meyer zu Schwabedissen HE, Kroemer HK (2011) In vitro and in vivo evidence for the importance of breast cancer resistance protein transporters (BCRP/MXR/ABCP/ABCG2). Handb Exp Pharmacol 325–371. Article Pubmed
[54] Dohse M, Scharenberg C, Shukla S, Robey RW, Volkmann T, et al. (2010) Comparison of ATP-binding cassette transporter interactions with the tyrosine kinase inhibitors imatinib, nilotinib, and dasatinib. Drug Metab Dispos 38:1371–1380. Article Pubmed
[55] Breedveld P, Beijnen JH, Schellens JH (2006) Use of P-glycoprotein and BCRP inhibitors to improve oral bioavailability and CNS penetration of anticancer drugs. Trends Pharmacol Sci 27:17–24. Article Pubmed
[56] Lo WL, Yu CC, Chiou GY, Chen YW, Huang PI, et al. (2011) MicroRNA-200c attenuates tumour growth and metastasis of presumptive head and neck squamous cell carcinoma stem cells. J Pathol 223:482–495. Article Pubmed
[57] Yu CC, Chen YW, Chiou GY, Tsai LL, Huang PI, et al. (2011) MicroRNA let-7a represses chemoresistance and tumourigenicity in head and neck cancer via stem-like properties ablation. Oral Oncol 47:202–210. Article Pubmed
[58] Wilson GD, Marples B, Galoforo S, Geddes TJ, Thibodeau BJ, et al. (2012) Isolation and genomic characterization of stem cells in head and neck cancer. Head Neck doi: 10.1002/hed.23184. Article Pubmed
[59] Wu MJ, Jan CI, Tsay YG, Yu YH, Huang CY, et al. (2010) Elimination of head and neck cancer initiating cells through targeting glucose regulated protein78 signaling. Mol Cancer 9:1–16. Article Pubmed
[60] Maitra A, Arking DE, Shivapurkar N, Ikeda M, Stastny V, et al. (2005) Genomic alterations in cultured human embryonic stem cells. Nat Genet 37:1099–1103. Article Pubmed
[61] Gu Y, Kim KH, Ko D, Nakamura K, Yasunaga Y, et al. (2004) A telomerase-immortalized primary human prostate cancer clonal cell line with neoplastic phenotypes. Int J Oncol 25:1057–1064. Article Pubmed
[62] Yasunaga Y, Nakamura K, Ko D, Srivastava S, Moul JW, et al. (2001) A novel human cancer culture model for the study of prostate cancer. Oncogene 20:8036–8041. Article Pubmed
[63] Sasaki K, Ichikawa H, Takei S, No HS, Tomotsune D, et al. (2009) Hepatocyte differentiation from human ES cells using the simple embryoid body formation method and the staged-additional cocktail. Scientific World Journal 9:884–890. Article Pubmed
[64] Chen SF, Nieh S, Jao SW, Liu CL, Wu CH, et al. (2012) Quercetin suppresses drug-resistant spheres via the p38 MAPK-Hsp27 apoptotic pathway in oral cancer cells. PLoS One 7:e49275 doi: 10.1371/journal.pone.0049275. Article Pubmed
[65] Tsai LL, Yu CC, Chang YC, Yu CH, Chou MY (2011) Markedly increased Oct4 and Nanog expression correlates with cisplatin resistance in oral squamous cell carcinoma. J Oral Pathol Med 40:621–628. Article Pubmed
[66] Gupta PB, Chaffer CL, Weinberg RA (2009) Cancer stem cells: mirage or reality? Nat Med 15:1010–1012. Article Pubmed
[67] Liu C, Tang D (2011) MicroRNA regulation of cancer stem cells. Cancer Res 71:5950–5954. Article Pubmed
[68] Dahler AL, Jones SJ, Dicker AJ, Saunders NA (1998) Keratinocyte growth arrest is associated with activation of a transcriptional repressor element in the human cdk1 promoter. J Cell Physiol 177:474–482. Article Pubmed
[69] Tsai WC, Lin YC, Tsai ST, Shen WH, Chao TL, et al. (2011) Lack of modulatory function of coding nucleotide polymorphism S100A2_185G>A in oral squamous cell carcinoma. Oral Dis 17:283–290. Article Pubmed
[70] Qiu W, Schönleben F, Li X, Su GH (2007) Disruption of transforming growth factor beta-Smad signaling pathway in head and neck squamous cell carcinoma as evidenced by mutations of SMAD2 and SMAD4. Cancer Lett 245:163–170. Article Pubmed
[71] Chang X, Monitto CL, Demokan S, Kim MS, Chang SS, et al. (2010) Identification of hypermethylated genes associated with cisplatin resistance in human cancers. Cancer Res 70:2870–2879. Article Pubmed



