Journal of Cancer Research & Therapy
An International Peer-Reviewed Open Access Journal
ISSN 2052-4994


- Download PDF
- |
- Download Citation
- |
- Email a Colleague
- |
- Share:
-
- Tweet
-

Journal of Cancer Research & Therapy
Volume 1, Issue 9, November 2013, Pages 208-214
Original researchOpen Access
Clinicopathological study of gene rearrangement and immunohistochemical pattern of primary intracranial diffuse large B-cell lymphomas
- 1 Department of Laboratory medicine, 456th Hospital of Jinan Military Command, Jinan 250031, China
- 2 Department of Laboratory medicine, 91th Hospital of Jinan Military Command, Jiaozuo 454003, China
- 3 Department of Nephrology, 456th Hospital of Jinan Military Command, Jinan 250031, Chian
- 4 Department of Laboratory medicine, 1st Sanatorium of Jinan Military Command, Qingdao 266071, China
*Corresponding author: Dr. Lihua Yu, Department of Laboratory medicine, 1st Sanatorium of Jinan Military command, Qingdao, China, 266071, Tel.: +86 532 51885559; Fax: +86 532 51885559. E-mail: pathology90@126.com
Han X and Xue C contributed equally to this study
Received 21 July 2013 Revised 10 October 2013 Accepted 19 October 2013 Published 27 October 2013
DOI: http://dx.doi.org/10.14312/2052-4994.2013-31
Copyright: © 2013 Han X, et al. Published by NobleResearch Publishers. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
AbstractTop
We studied the clinicopathological and imaging characteristics of primary intracranial diffuse large B-cell lymphomas (PIC-DLBCL). Imaging, histopathological findings, and immunohistochemical staining characteristics were analyzed, and the immunoglobulin heavy and light chain gene rearrangement of 25 PIC-DLBCL cases was examined. MicroRNA was extracted from 10 cases each of PIC-DLBCL, extracerebral germinal center DLBCL (GC-DLBCL), and extracerebral non-GC-DLBCL (NGC-DLBCL); we conducted chip hybridization and comparatively analyzed the difference among the three. PIC-DLBCLs typically involved no less than two cerebral lobes (10/25); the frontal lobe was affected most often (6/25). Target-shaped structures were observed in all PIC-DLBCLs due to the proliferation of centroblast-like large lymphocytes surrounding the vessels. There was strong and diffuse immunostaining for CD20 and CD79a, and negative immunostaining for CD3, CD5, CD23, and cyclin D1 for all PIC-DLBCLs. The percentage of cells with nuclear positivity for anti-Ki67 antibody ranged 50-90% (mean, 80%). Three, 19, and 22 PIC-DLBCLs were CD10-, Bcl-6-, and melanoma ubiquitous mutated 1-positive, respectively. Twenty-four PIC-DLBCLs were B-cell monoclonal. MicroRNA hybridization showed that 788 PIC-DLBCL microRNAs/segments increased to at least twice of that of NGC-DLBCLs, and 401 PIC-DLBCL microRNAs/segments declined to less than half of that of NGC-DLBCLs. Six hundred and eleven PIC-DLBCL microRNAs/segments increased to at least twice that of GC-DLBCLs, and 229 PIC-DLBCL microRNAs/segments declined to less than half of that in GC-DLBCLs. PIC-DLBCL typically affected multiple sites, tended to occur in older men, arose from activated B cells, had high B-cell monoclonality; its microRNA expression differed from that of NGC-DLBCL and GC-DLBCL.
Keywords: intracranial; diffuse large B-cell lymphoma; gene rearrangement; microRNA
IntroductionTop
Primary intracranial lymphoma (PICL) is very rare, early diagnosis is difficult, and prognosis is poor; it accounts for 0.8-1.5% of primary intracranial tumors [1, 2], while primary intracranial lymphoma diffuse large B-cell lymphoma (PIC-DLBCL) accounts for 95-98% of PICL. As there is no lymphatic circulation in the central nervous system, the origin and mechanism of neoplasia of lymphoma cells are unknown. We studied and analyzed the characteristics of PIC-DLBCL through imaging, immunohistochemistry, gene rearrangement detection, and microRNA expression to discuss the clinicopathological characteristics of PIC-DLBCL and contribute to the understanding of its clinical character, tissue origin, diagnosis and prognosis.
Materials and methodsTop
Case data
We studied 25 cases of PIC-DLBCLs that had been admitted and diagnosed by Laboratory medicine of 456th Hospital and 91th Hospital of Jinan Military command over 11 years from January 2000 to December 2011. 15 cases were male and 10 were female; ages ranged 19-78 years (mean, 59.64 years); 10 cases had involvement of two or more lobes; 6 cases involved only the frontal lobe; 9 cases involved other lobes. No case had hereditary or acquired immunodeficiency. Each case was accompanied by imaging data and underwent immunohistochemical staining and detection of B-cell immunoglobulin (Ig) heavy and light chain gene rearrangement. Of these, 10 cases expressing Bcl-6 and melanoma ubiquitous mutated 1 (Mum-1) were selected for microRNA chip hybridization detection; 10 cases of peripheral germinal center DLBCL (GC-DLBCL) and 10 cases of non-GC-DLBCL (NGC-DLBCL) were selected for microRNA chip hybridization for comparison. All patients gave informed consent, and the trial was approved by the Medical Ethical Committee of this two hospital.
Antibodies
Rat anti-human primary antibodies to CD20, CD79a, CD3, CD5, CD23, cyclin D1, CD10, and Bcl-6, rabbit anti-human Ki67 antibody, secondary antibody, and diaminobenzidine color-developing agent were purchased from Santa Cruz Biotechnology, while rabbit anti-human primary antibody to Mum-1 was purchased from Novocastra.
Kits
A TIANamp Genomic DNA Kit for DNA purification was purchased from TianGen Biotech (China). An immunoglobulin heavy and light chain gene rearrangement detection kit was purchased from InVivoScribe Technologies. MicroRNA chip detection was carried out using a miRCURY LNA™ microRNA Array V.14.0 and a miRCURY™ Array Power Labeling kit purchased from Danish Exiqon.
Experimental method
After conventional hematoxylin-eosin (HE) staining, the histopathological characteristics of sections obtained from each PIC-DLBCL case were observed and analyzed.
Immunohistochemical staining
Paraffin sections (4-µm thick) from each PIC-DLBCL case were adhered on glass slides, treated with 3-aminopropyltriethoxysilane, and stained using the Ventana BenchMark XT system (Roche Ltd.).
Detection of heavy and light chain gene rearrangement
DNA extraction
Each paraffin-embedded tissue slice (6 µm) was dewaxed three times with dimethylbenzene, and then washed three times with absolute ethyl alcohol. After swabbing, DNA was extracted and labeled according to the manufacturer’s instructions, and then stored at 4°C before using.
PCR
A 20-µL reaction system containing 18 µL PCR mix, 0.1 µL Taq enzyme, 2 µL template DNA was used. The PCR reaction program was as follows: denaturation 45 seconds at 95°C, annealing 45 seconds at 60°C, extension 90 seconds at 72°C for 34 cycles.
Electrophoresis
Electrophoresis was carried out on 8% polyacrylamide gel at 80 V for 1 h.
Identification of results
DNA bands were observed in IgH tube A/B/C/D/E, IgK tube A/B, and IgL tube; an internal control was used. One or two bands with sharp edges (1-mm wide, without trails or smears) were identified as monoclonal as they were within the corresponding molecular weight.
MicroRNA chip hybridization
Using HE-stained slices as contrast, paraffin-embedded tissue slices from 10 typical PIC-DLBCL, GC-DLBCL, and NGC-DLBCL cases each were microdissected with a tissue chip device, and then chip hybridization was carried out using the microRNA extracted from the microdissected tissues. The tissue chip device used to perform the tumor tissue microdissection involved a 2-mm bodkin that was used to drill a 3-mm strip of tissue from each case to test the chip expression pattern.
Total RNA in the microdissected tumor cells was extracted with the TRIzol one-step method. The microRNA from each specimen was labeled using the miRCURY™ Array Power Labeling kit. Specimen hybridization solution (180 µL) was prepared according to the kit instructions. The hybridization signals were scanned into a computer with an Axon GenePix 4000B microchip scanner. The original densities of the acquired signals were read by GenePix pro V6.0 software and the data were analyzed and compared.
Statistical analysis
Statistical analysis was carried out with SPSS 10.0 software; relevance was tested with the Fisher’s precision parameter.
ResultsTop
Imaging
Magnetic resonance imaging (MRI) revealed decreased signal intensity on T1-weighted imaging (T1WI), equal or slightly increased signal intensity on T2-weighted imaging (T2WI), and increased signal intensity (edema region) around the tumor of the 25 PIC-DLBCL cases. The intensity was well distributed in solid-appearing tumors (Figure 1).
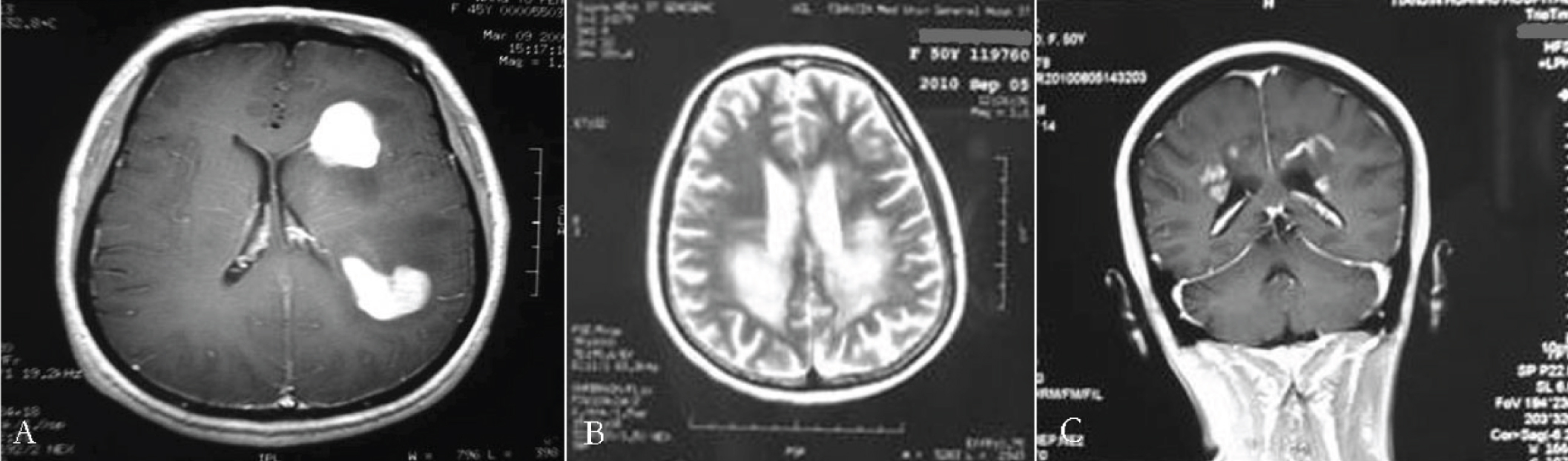
Figure 1 (A) MRI well-distributed intensity on a solid-appearing multiple tumor (intensity based on T1 weighting); (B) Multi-part MRI T2 weighting; (C) Multi-part MRI intensity imaging.
Pathological histological characteristics
In all cases, light microscopy revealed centroblast-like large lymphocyte proliferation that surrounded the vessels. Tumor cells exhibited significant atypia. However, glioblastoma multiforme and glomerulus-like proliferating vascular endothelial cells were absent; clear, flake-shaped necrosis was not observed. Adhesion between the tumor cells was poor, and there were nucleoli of irregular size and sporadic infiltration of small lymphocytes between the tumor cells (Figure 2A, 2B, 2C).
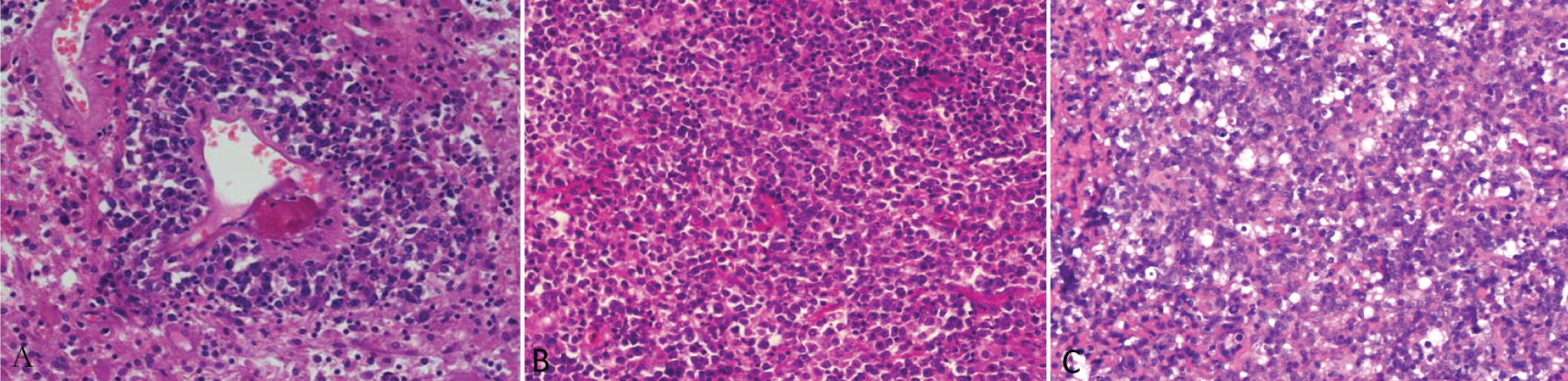
Figure 2 (A) Lymphoma cells’ scoring-ring shape growth around the blood vessels; (B) Tumor cells’ diffuse and invasive growth; (C) Tumor cells’ diffuse and invasive growth with significant atypia.
Immunohistochemistry
In all 25 PIC-DLBCL cases, there was diffuse, strong immunostaining for CD20 and CD79a, and negative immunostaining for CD3, CD5, CD23, and cyclin D1. The percentage of cells with nuclear positivity for anti-Ki67 antibody ranged 50-90% (mean, 80%). Three, 19, and 22 cases expressed CD10 (12%), Bcl-6 (76%), and Mum-1 (88%), respectively. Apart from tumor cells, there were sporadic CD3- and CD20-positive small lymphocytes (Figure 3A, 3B, 3C, 3D).
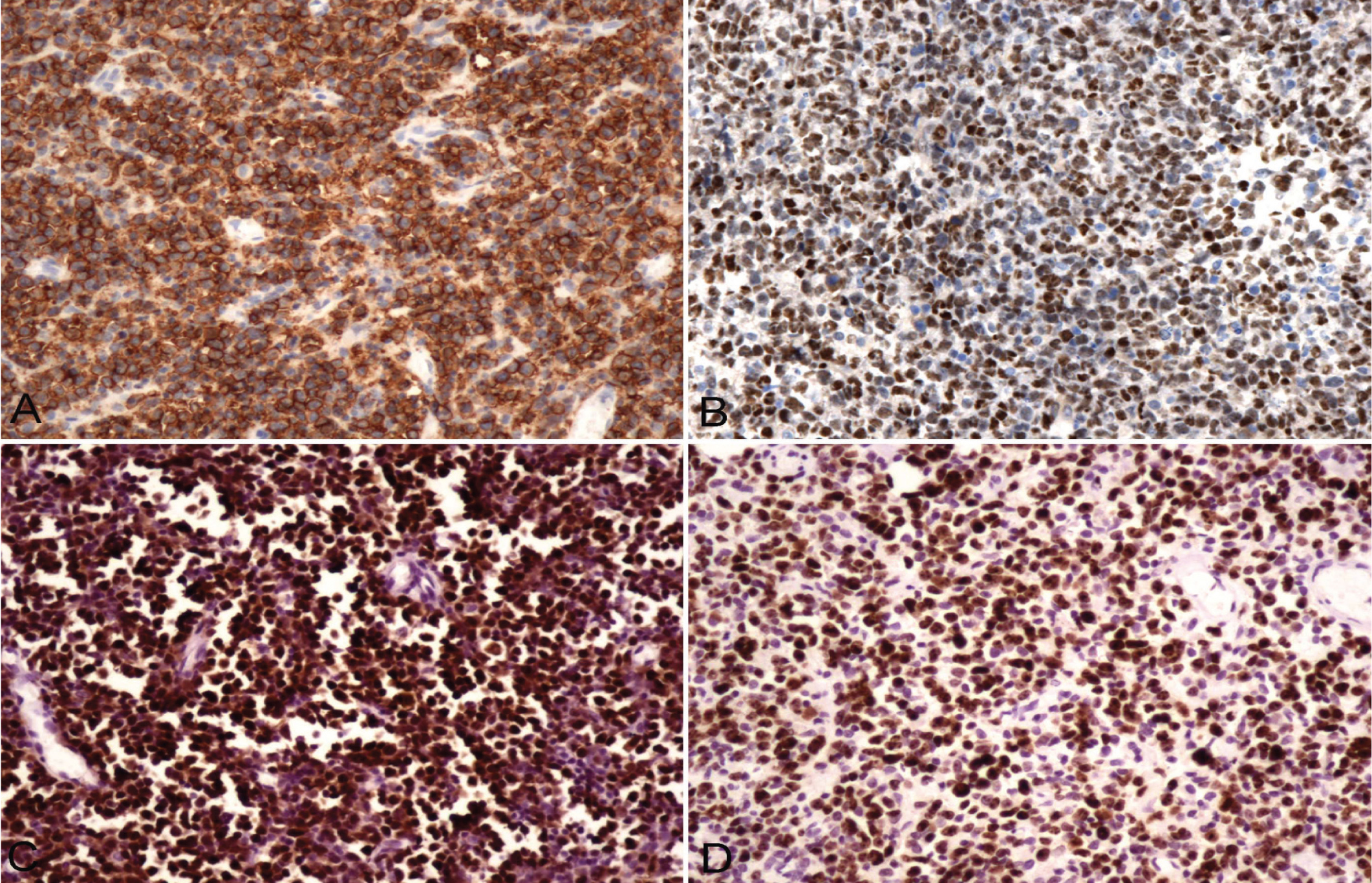
Figure 3 A) Tumor cells were diffuse and strongly positively express CD20 through EnVisin method; (B) Tumor cells were diffuse and strongly positively express bcl-6 through EnVisin method; (C) Tumor cells expressed Mum-1 through EnVisin method; (D) Tumor cells expressed Ki-67 at high proportion through EnVisin method.
Gene rearrangement
Of 25 PIC-DLBCLs, 24 (96%) were B-cell monoclonal; of these, 16 were heavy chain monoclonal, 11 were light chain monoclonal, and 10 were both heavy and light chain-positive (Figure 4).
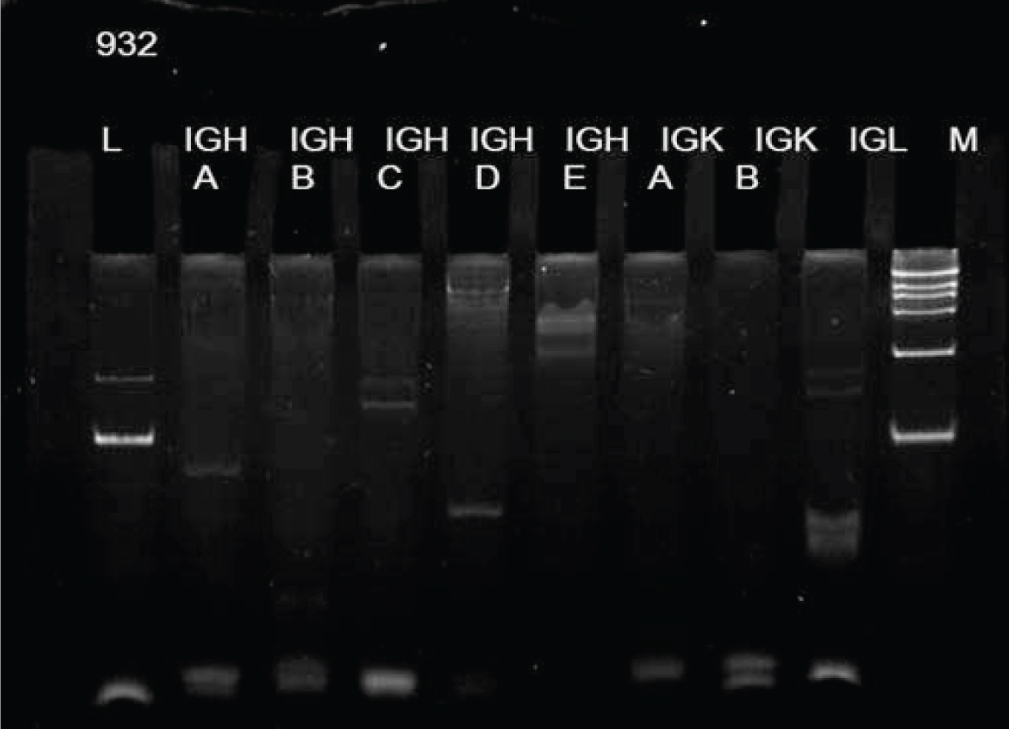
Figure 4 lymphocyte gene rearrangement detection showed heavy chain and light chain were both monoclonal.
MicroRNA relevance between specimens
The microRNA relevance of PIC-DLBCL to GC-DLBCL (0.7608) was higher than that of PIC-DLBCL to NGC-DLBCL (0.6440), and the relevance of GC-DLBCL to NGC-DLBCL was the highest: 0.8794 (Table 1).
| Lymphoma type | GC-DLBCL | PIC-DLBCL |
| GC-DLBCL | 1 |
0.7608 |
| PIC- DLBCL | 0.7608 |
1 |
| NGC-DLBCL | 0.8794 |
0.644 |
MicroRNA chip hybridization
Compared with NGC-DLBCL, microRNA chip hybridization (Figure 5) showed that 28 microRNA or segments for PIC-DLBCL cases (Table 2) increased by 10 times or more, while 788 microRNA or segments of PIC-DLBCL increased by two times or more; 401 microRNA or segments of PIC-DLBCL decreased to 0.5 times or more, while 33 microRNA or segments decreased to 0.1 times or more (Table 3). Compared with GC-DLBCL, 11 microRNA or segments (Table 4) increased by 10 times or more, while 611 microRNA or segments of PIC-DLBCL increased by two times or more; 229 microRNA or segments of PIC-DLBCL decreased to 0.5 times or more, while 13 microRNA or segments (Table 5) decreased to 0.1 times or more.
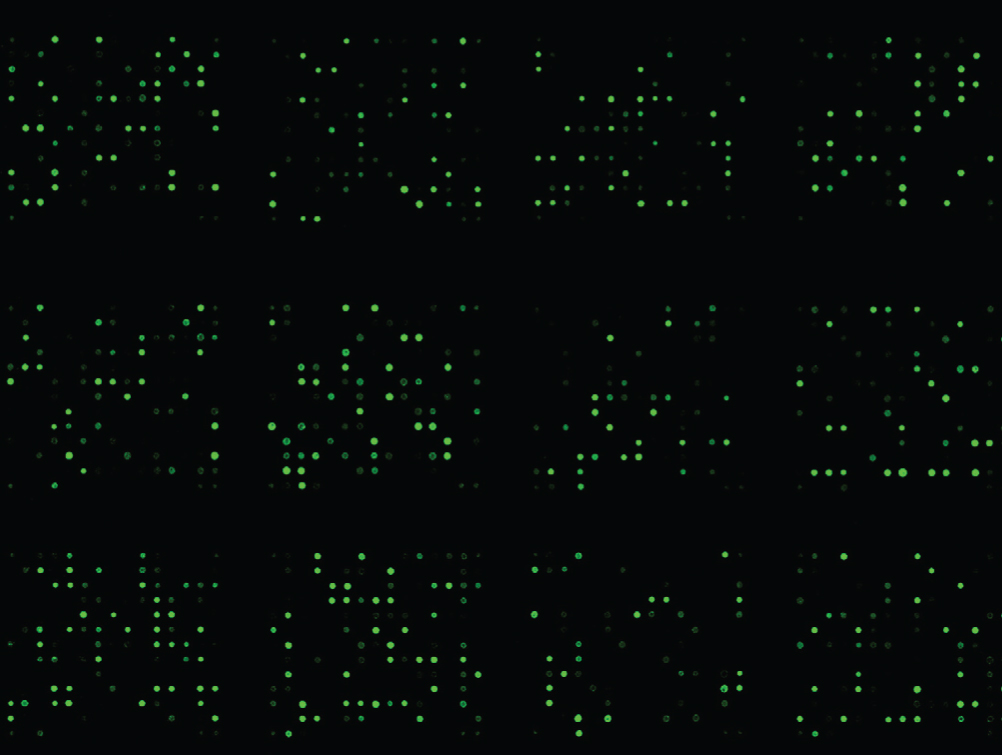

Figure 5 PIC- DLBCLmicroRNA chip hybridization (partial).
| ID | microRNA | PIC- DLBCL / NGC -DLBCL |
| 19583 | hsa-miR-122 |
25.60826 |
| 17540 | 18.06316 |
|
| 11255 | 17.69825 |
|
| 146031 | 15.87368 |
|
| 42779 | 15.43579 |
|
| 17525 | 14.23158 |
|
| 28581 | spike_control_v2_36 |
14.01263 |
| 42880 | 13.41952 |
|
| 46789 | hsa-miR-513b |
13.31231 |
| 46804 | hsa-miRPlus-E1027 |
12.80842 |
| 42714 | 12.58947 |
|
| 46885 | hsa-miRPlus-E1225 |
12.58947 |
| 46641 | hsa-miRPlus-E1101 |
12.5274 |
| 46370 | hsa-miRPlus-E1150 |
12.51128 |
| 17668 | hsa-miR-552 |
12.49288 |
| 46197 | SNORA66 |
12.45263 |
| 46322 | 12.04211 |
|
| 42878 | 11.98649 |
|
| 33116 | 11.63468 |
|
| 146059 | 11.49474 |
|
| 146165 | hsa-miR-1973 |
11.13671 |
| 42631 | 10.99713 |
|
| 42569 | 10.94737 |
|
| 17388 | 10.84785 |
|
| 146060 | 10.62476 |
|
| 145721 | hsa-miR-875-5p |
10.4 |
| 42581 | hsa-miR-513a-5p |
10.30353 |
| 146157 | hsv1-miR-H4 |
10.12632 |
| ID | microRNA | PIC- DLBCL / NGC -DLBCL |
| 145720 | hsa-miR-454 |
0.098879 |
| 27533 | hsa-miR-320a |
0.098698 |
| 145741 | hsa-miR-545 |
0.09731 |
| 10988 | hsa-miR-194 |
0.09707 |
| 10952 | hsa-miR-146a |
0.096997 |
| 17814 | hsa-miR-570 |
0.095195 |
| 42965 | hsa-miR-424 |
0.09489 |
| 145947 | hsa-miRPlus-C1075 |
0.091228 |
| 42496 | hsa-miR-181c |
0.088285 |
| 17608 | hsa-miR-425 |
0.084909 |
| 46803 | hsa-miR-503 |
0.082105 |
| 10995 | hsa-miR-199a-3p/hsa-miR-199b-3p |
0.081434 |
| 19591 | hsa-miR-199b-5p |
0.081092 |
| 42630 | hsa-miR-140-3p |
0.075453 |
| 42839 | hsa-miR-135a |
0.073969 |
| 4700 | hsa-miR-140-5p |
0.072543 |
| 42847 | hsa-miR-497 |
0.071233 |
| 29562 | hsa-miR-199a-5p |
0.069304 |
| 11078 | hsa-miR-365 |
0.06884 |
| 145844 | hsa-miR-374a |
0.067405 |
| 146206 | 0.06239 |
|
| 11015 | hsa-miR-215 |
0.060569 |
| 11024 | hsa-miR-223 |
0.056999 |
| 42912 | hsa-miR-339-3p |
0.055081 |
| 11018 | hsa-miR-218 |
0.049202 |
| 10916 | hsa-miR-1 |
0.04812 |
| 10942 | 0.039705 |
|
| 10987 | hsa-miR-193b |
0.038129 |
| 17946 | hsa-miR-192* |
0.03775 |
| 10925 | hsa-miR-10b |
0.034847 |
| 17732 | hsa-miR-192 |
0.033495 |
| 10990 | hsa-miR-196a |
0.020655 |
| 11077 | hsa-miR-363 |
0.01578 |
| ID | microRNA | PIC- DLBCL / GC -DLBCL |
| 46789 | hsa-miR-513b |
19.96699 |
| 17525 | 15.09825 |
|
| 146157 | hsv1-miR-H4 |
14.32398 |
| 11255 | 14.08202 |
|
| 45891 | hsa-miRPlus-E1285 |
13.63466 |
| 146041 | 12.77544 |
|
| 14294 | 11.90439 |
|
| 146177 | 11.06749 |
|
| 30442 | hsa-miR-802 |
10.45263 |
| 11144 | hsa-miR-512-3p |
10.45263 |
| 46256 | hsa-miRPlus-E1238 |
10.1824 |
| ID | microRNA | PIC- DLBCL / GC -DLBCL |
| 10925 | hsa-miR-10b |
0.094633 |
| 46842 | hsa-miRPlus-E1194 |
0.089339 |
| 145755 | hsa-miR-146a* |
0.084224 |
| 145947 | hsa-miRPlus-C1075 |
0.080097 |
| 42912 | hsa-miR-339-3p |
0.072025 |
| 19591 | hsa-miR-199b-5p |
0.071811 |
| 29328 | hsa-miR-582-3p |
0.064522 |
| 42800 | 0.064522 |
|
| 10942 | 0.054931 |
|
| 42544 | hsa-miR-582-5p |
0.047297 |
| 145679 | 0.041906 |
|
| 17312 | hsa-miR-592 |
0.013965 |
| 42839 | hsa-miR-135a |
0.013584 |
DiscussionTop
PICL morbidity has increased significantly in recent years. Its incidence among other primary brain tumors is reported to have increased from 0.8-1.5% to 6.6% [3]. More than 95% of CSNLs can be DLBCL. Early diagnosis of PIC-DLBCL is difficult: due to its complex image expression and atypical clinical symptoms, it is often misdiagnosed as glioma, metastatic tumor, or inflammation. This study found that PIC-DLBCL was prevalent in elderly males and with multiple sites of involvement. Plain computed tomography indicated that lesions were quasicircular or irregularly shaped; 11 cases were low density or equidensity, while 14 were high density. MRI revealed decreased signal intensity on T1WI, equal or slightly increased signal intensity on T2WI, and increased signal intensity (edema region) around the tumor. Blood, calcification, or cystic degeneration was not found in the lesions. In addition, signal intensities for all cases increased significantly after enhancement and most were lump-shaped; “gap-shaped” intensification was observed for five cases, which provided a theoretical basis for early diagnosis and differential diagnosis of PIC-DLBCL. The cell-origin and pathogenesis of PIC-DLBCL is still being debated because there is no lymphatic circulation in the central nervous system.
From the pathological histology viewpoint, the tumor cells exhibited diffuse and invasive growth, tending to infiltrate small brain vessels and gather in the peripheral spaces of vascular walls, forming “oversleeve” structures and perivascular cuffing of the tumor cells. Moreover, the cortex or nerve tract inside the white matter infiltrated by tumor cells was visible. Necrosis might have been present inside the tumor tissues, accompanied by surrounding small lymphocyte infiltration. Occasionally, large reactive astrocytes were observed between tumors or their peripheral tissues. PIC-DLBCL is mostly composed of centroblast-like lymphocytes; part of the nucleolus is noticeable and similar to an immunoblast. The morphology of the tumor cells was almost identical to that of extracerebral DLBCL. However, there is a significant difference clinically between the survival rates of PIC-DLBCL and extracerebral DLBCL.
In recent years, researchers have been able to divide extracerebral DLBCL into GC and NGC origin [4, 5] by immunohistochemical staining. Tumor cells that express CD10 or coexpress CD10 and Bcl-6 rather than Mum-1 are classified into the GC origin group, while tumor cells expressing Mum-1 or coexpressing Bcl-6 and Mum-1 are classified into the NGC origin group. Our immunohistochemical staining results demonstrated that only three and 22 cases could be classified into the GC and NGC origin group, respectively, in accordance with the literature [6]. In addition, one feature of centroblast-like tumor cells expressing CD20 was growth surrounding the vessels, and there were sporadic CD3- and CD20-positive small lymphocytes. This indicated that tumor cells were malignantly transformed against a background of T/B lymphocyte mixed inflammation. Moreover, lymphocyte malignant transformation mainly involved polyclonal to oligoclonal and polyclonal to monoclonal hyperplasia. Monoclonal detection for lymphoid tissues can aid in lymphoma diagnosis and understanding of the mechanism of tumor development. The present study found that the monoclonal immunoglobulin-positive rate of each B-cell lymphoma subtype was 94-100%, and a 99% total positive rate has been reported [7]. In this study, 96% of cases were B-cell monoclonal, which indicates that PIC-DLBCL and DLBCL are also monoclonal. The morphology and clonality of PIC-DLBCL were similar to that of extracerebral DLBCL; however, whether pathogenesis is identical between these two groups requires further study.
MicroRNA plays an important role in tumorigenesis and can be used for accurate tumor classification and evaluation of prognosis [8]. In recent years, microRNA has also been used in the study of DLBCL [9, 10]. Malumbres et al. [11] reported that there were specific microRNA expression at different B-cell differentiation stages. They indicated in a further study that nine microRNAs could distinguish DLBCL into two major subtypes. Using miRBase V12.0, Culpin et al. [12] tested microRNA chips of GC- and activated cell-origin DLBCL, and found nine and four diverse microRNAs between DLBCL and follicular lymphoma, respectively. Baraniskin et al. [13] reported that the level of microRNA in cerebrospinal fluid could aid in the diagnosis of PICL. A tissue chip device has been introduced for sectioning tumor tissues, and the present study used miRBase V14.0 for the microRNA chip hybridization. We found that the microRNA expression of PIC-DLBCL was significantly different from that of NGC-DLBCL and GC-DLBCL. NGC-DLBCL and GC-DLBCL had the highest microRNA relevance, followed by PIC-DLBCL and GC-DLBCL; the relevance was lowest between PIC-DLBCL and NGC-DLBCL. From these results, it is clear that PIC-DLBCL is significantly different from extracerebral DLBCL, which at least clarifies that PIC-DLBCL does not originate directly from extracerebral B lymphocytes; it is likely that it develops from malignant transformation of polyclonal B lymphocytes in the intracralvarium via monoclonal hyperplasia.
ConclusionTop
PIC-DLBCL is a relatively malignant heterogeneous tumor with quick growth, short disease course, and poor prognosis. However, it is sensitive to cortical hormone therapy, radiotherapy, and chemotherapy. Overall survival for PIC-DLBCL can be expected to be extended following familiarity with its imaging and clinicopathological characteristics, increased understanding, accurate early diagnosis, and multidisciplinary treatment modalities with high-dose methotrexate as basic medication; however, its prognostic indicator, causes, and pathogenesis require further investigation.
Conflict of interest
The authors wish to express that they have no conflict of interest.
ReferencesTop
[1] Pasricha S, Gupta A, Gawande J, Trivedi P, Patel D (2011) Primary central nervous system lymphoma: a study of clinicopathological features and trend in western India. Indian J Cance 48:199-203. Article Pubmed
[2] Bhagavathi S, Wilson JD (2008) Primary central nervous system lymphoma. Arch Pathol Lab Med 132:1830-1834. Article Pubmed
[3] L Z. Diagnosis and Treatment Progress of PCNSL. Minim Invas Neurosurg 2008; 13: 186-189.
[4] Anderson JJ, Fordham S, Overman L, Dignum H, Wood K (2009) Immunophenotyping of diffuse large B-cell lymphoma (DLBCL) defines multiple sub-groups of germinal centre-like tumours displaying different survival characteristics. Int J Oncol 35: 961-971. Article Pubmed
[5] Y W, J L, C Z (2012) Analysis of clinicopathologic characteristics and immunophenotype of 16 cases of PCNSL. J Clin Exp Pathol 28:1322-1328.
[6] Hattab EM, Martin SE, Al-Khatib SM, Kupsky WJ, Vance GH, et al. (2010) Most primary central nervous system diffuse large B-cell lymphomas occurring in immunocompetent individuals belong to the nongerminal center subtype: a retrospective analysis of 31 cases. Mod Pathol 23:235-243. Article Pubmed
[7] Evans PA, Pott Ch, Groenen PJ, Salles G, Davi F, et al. (2007) Significantly improved PCR-based clonality testing in B-cell malignancies by use of multiple immunoglobulin gene targets. Report of the BIOMED-2 Concerted Action BHM4-CT98-3936. Leukemia 21:207-214. Article Pubmed
[8] Doleshal M, Magotra AA, Choudhury B, Cannon BD, Labourier E, et al. (2008) Evaluation and validation of total RNA extraction methods for microRNA expression analyses in formalin-fixed, paraffin-embedded tissues. J Mol Diagn 10:203-211. Article Pubmed
[9] Nie K, Zhang T, Allawi H, Gomez M, Liu Y, et al. (2010) Epigenetic down-regulation of the tumor suppressor gene PRDM1/Blimp-1 in diffuse large B cell lymphomas: a potential role of the microRNA let-7. Am J Pathol 177:1470-1479. Article Pubmed
[10] Robertus JL, Harms G, Blokzijl T, Booman M, de Jong D, et al. (2009) Specific expression of miR-17-5p and miR-127 in testicular and central nervous system diffuse large B-cell lymphoma. Mod Pathol 22:547-555. Article Pubmed
[11] Malumbres R, Sarosiek KA, Cubedo E, Ruiz JW, Jiang X, et al. (2009) Differentiation stage-specific expression of microRNAs in B lymphocytes and diffuse large B-cell lymphomas. Blood 113:3754-3764. Article Pubmed
[12] Culpin RE, Proctor SJ, Angus B, Crosier S, Anderson JJ, et al. (2010) A 9 series microRNA signature differentiates between germinal centre and activated B-cell-like diffuse large B-cell lymphoma cell lines. Int J Oncol 37:367-376. Article Pubmed
[13] Baraniskin A, Kuhnhenn J, Schlegel U, Chan A, Deckert M, et al. (2011) Identification of microRNAs in the cerebrospinal fluid as marker for primary diffuse large B-cell lymphoma of the central nervous system. Blood 117:3140-3146. Article Pubmed



