Journal of Cancer Research & Therapy
An International Peer-Reviewed Open Access Journal
ISSN 2052-4994


- Download PDF
- |
- Download Citation
- |
- Email a Colleague
- |
- Share:
-
- Tweet
-

Journal of Cancer Research & Therapy
Volume 1, Issue 1, March 2013, Pages 11–23
Original researchOpen Access
Sonic hedgehog inhibition reduces in vitro tumorigenesis and alters expression of Gli1-target genes in a desmoplastic medulloblastoma cell line
-
García-López R1,
Vera-Cano B1,
Vacas-Oleas A1,
de la Rosa J1,
Gallo-Oller G1,
Alonso MM2,
Rey JA3 and
Castresana JS1,*

- 1 Brain Tumor Biology Unit, University of Navarra School of Sciences, 31008 Pamplona, Spain
- 2 Department of Oncology, Clínica Universidad de Navarra, 31008 Pamplona, Spain
- 3 Unidad de Investigación, IdiPAZ, Hospital Universitario La Paz, 28046 Madrid, Spain
*Corresponding author: Castresana JS, Unidad de Biología de Tumores Cerebrales, Edificio de Investigación, 2261, Universidad de Navarra, Irunlarrea 1, 31008 Pamplona, Spain. Tel.: +34 948 425600 (ext. 806486); Fax: +34 948 425740; E-mail: jscastresana@unav.es. E-mail: jscastresana@unav.es
Received 2 November 2012 Revised 5 January 2013 Accepted 13 January 2013 Published 20 January 2013
DOI: http://dx.doi.org/10.14312/2052-4994.2013-3
Copyright: © 2013 García-López R, et al. Published by NobleResearch Publishers. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
AbstractTop
Medulloblastoma is one of the most frequent and aggressive tumors of childhood. The Sonic hedgehog (Shh) pathway, related to human development, is altered in most medulloblastomas: genes like Ptch, Smo, or Sufu suffer mutations in 15% to 25% of these tumors. We tested Shh inhibition in the Daoy medulloblastoma cell line by two methods: a molecular one, direct Gli1 siRNA inhibition; and a pharmacological inhibition of Smo, upstream of Gli1, by cyclopamine. Afterwards, a comparison of cellular and molecular responses was done. In general, we proved that cell viability, cell migration and cell colony formation decreased after Shh inhibition, which might confer a less tumorigenic status to Daoy cells. Moreover, we assessed the expression of different Gli1 target genes and other genes and found that Shh shows a crosstalk with oncogenes and tumor suppressor genes that have been described in numerous tumors. All these experiments give an overview of the Shh pathway in medulloblastoma, together with the demonstration of the efficacy of cyclopamine and Gli1 siRNA Shh inhibition in vitro.
Keywords: sonic hedgehog; cyclopamine; siRNA; Gli1, Ptch; Smo; in vitro tumorigenesis; medulloblastoma; Dao
IntroductionTop
Medulloblastoma (MB) is the most common tumor of the posterior fossa in childhood, representing 30% to 40% of tumors in this age group and one of the most common pediatric solid tumors (15% to 25%) with an annual incidence of about 6.5 cases per million population in Europe [1] . Incidence is significantly higher in boys than in girls (approximately 60% in boys). Although its world incidence is lower than in other tumors, the treatment of these tumors that includes surgical resection, craniospinal radiation and chemotherapy only cures 60% of the affected children and most suffer from long-term side effects derived from the aggressiveness of the treatment [2 , 3]. Due to limitations of the current treatment, the understanding of signaling pathways involved in medulloblastoma pathogenesis might lead to the design of new targeted treatments against medulloblastoma.
The most important signaling pathways that correlate with medulloblastoma development are those that regulate the formation of granule cell neurons and neural stem and precursor cell proliferation in the central nervous system (CNS), allowing a medulloblastoma classification that could also serve as a guide for successful anti-neoplastic therapy. According to the current consensus nomenclature [4] , Wingless (Wnt) signaling pathway gives the name to a subgroup of medulloblastomas, as does the Sonic hedgehog (Shh) pathway. Alterations in both affect 30% to 40% of medulloblastomas [5 , 6]. Mutations activating the Wnt pathway member beta-catenin are more common than adenomatous polyposis coli (APC) loss in sporadic medulloblastoma and are found in 5% to 10% of cases [7, 8]. The two remaining subgroups of medulloblastoma classification (groups 3 and 4) are based on a clustering algorithm [9] and are different in terms of demographics, histology, DNA copy-number aberrations and clinical outcome [10]. A number of other genes important in cerebellar stem and progenitor cells are deregulated in medulloblastoma. These genes include the oncogene N-myc, Bmi-1 and Otx2, bone morphogenic proteins (Bmp), the Notch cell signaling pathway and others.
The Shh signaling pathway is a major mitogenic regulator of granule neuron precursor cells (GNPCs) [11] . The mitogen Sonic hedgehog (Shh) drives proliferation of GNPCs by binding to the 12 transmembrane receptor Patched (Ptch) [12] . In the absence of Shh, Ptch represses the function of Smoothened (Smo), a seven transmembrane G-protein-coupled receptor-like protein that activates the Gli1 and Gli2 transcription factors and inactivates the transcriptional repressor Gli3 that together regulate the transcriptional program in the cell nucleus triggering transcription of several Shh target genes including Gli1, Ptch1, Hh interacting protein (Hhip), Cyclin D2, N-Myc, Bmi-1 and Bcl-2 [11, 13]. In addition, transfection of Gli1 results in decreased expression of Plakoglobin, Embigin and TSC-22 [14] . The transduction cascade from Smo to Gli is regulated at several different levels by activators such as Dyrk1, IFTs, Ras and Akt and inhibitors such as Sufu, Ren, PKA and GSK3β (Figure 1).
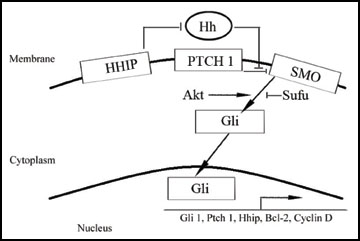
Figure 1 The sonic hedgehog pathway regulators.
Mutations in Shh pathway genes are found in approximately 25% of medulloblastomas [15] . Particularly, somatic mutations in the Ptch receptor, which result in activation of the Shh pathway, have been identified in 15% of cases. The Shh pathway genes Smo and Sufu, suppressor of fused (protein of Gli Drosophila analog complex), are also occasionally mutated in human medulloblastoma [16, 17].
Moreover, there are certain homeobox transcription factors such as NK2 homeobox 2 (Nkx2.2) and paired box gene 6 (Pax6) that are related to Shh. Nkx2.2 is induced by Shh signaling during neuronal development [18] , while Pax6 is inhibited by Shh signaling during neuronal embryonic development. Other associated molecules are secreted frizzled-related protein 1 (SFRP1), downregulated by Shh signaling [19] , Bmi-1 that shows overexpression in human medulloblastoma [20] and hedgehog interacting protein (HHIP) that functions as an endogenous antagonist for Shh [21] .
Medulloblastoma associates with an inappropriate Shh-Gli activity [22] . Therefore, it may be sensitive to treatments with Shh pathway inhibitors such as cyclopamine, a plant-derived alkaloid that antagonizes this pathway by blocking Smo which triggers Gli to bind DNA and express its target genes [23] . From a genetic approach the pathway can be inhibited by siRNA against Gli1 resulting in a high and specific knockdown.
Therefore, our working hypothesis is that directly targeting Smo and Gli1 will enable us to understand better the core pathway its regulation and role in desmoplastic medulloblastomas.
In general, the aim of this work was to analyse the role of the Sonic hedgehog signaling pathway and its connections with other genes which regulate the proliferation and tumor development of medulloblastoma. An adhesive cell line as an initial study was used to be treated with down stream inhibitor (siRNA against Gli1) and also to prove the effect of a pharmacological treatment by an upstream antagonist (cyclopamine). In this sense, the two treatments were compared performing molecular and cellular trials using a desmoplastic medulloblastoma cell line, including assessment of cell proliferation, in vitro colony forming tumorigenic capacity, viability, and gene expression. All these experiments will give an overview of the Shh pathway in medulloblastoma, together with the possibility to demonstrate the efficacy of cyclopamine versus Gli1 siRNA and whether it could be a good strategy against human medulloblastoma in the near future.
Materials and methodsTop
Cell line
We used a desmoplastic medulloblastoma cell line: Daoy. This was purchased from
the American Type Culture Collection, Manassas, VA, USA. The cell line was cultured
in RPMI L-glutamax medium (Gibco-BRL, Gaithersburg MD, USA), supplemented with 10%
fetal bovine serum, 4% non-essential amino acids, 1% penicillin, and 0.1% amphotericin
B and maintained in an incubator at 37 oC
in an atmosphere with 5% CO2 to expand the
cells. Subculture of cells was performed after 80% confluence with the help of trypsin/EDTA
(Gibco).
RNA extraction and retrotranscription
RNA was extracted using the RNAeasy KIT (with DNAsa treatment included) (Qiagen)
following manufacturer’s instructions. Purity and quantity of total RNA was determined
by nanodrop measurement. Next, 2 µg RNA were retrotranscribed in a final volume
of 20 µL. RNA was preincubated at 72 oC
for 10 min with 250 µg random primers to eliminate secondary structures. After chilling
on ice, 1x RT-buffer synthesis buffer was added, 0.01 M DTT and 0.5 mM of each dNTP.
The reaction mixture was heated for 2 min at 42 oC
for random primer annealing. Then, 1 U™ II Reverse Transcriptase was added. cDNA
was synthesized at 42 oC for 40 min and
stored at -20 oC until use.
Quantitative RT-PCR
For qRT-PCR of HPRT, Gli1, Ptch1, Smo, Sufu, Gli3, cyclin D2, plakoglobin, Nkx2.2,
Bmi-1, Pax6, N-myc, Notch1 and Notch2, the amplification reactions were carried
out in an IQ5 multicolor real-time PCR detection system (BioRad, Hercules, CA, USA).
75 ng of template DNA were used in a total volume of 25 µL, the reaction mix contained
12.5 µL 2X IQ™ SYBR Green Supermix and 12.5 pmol forward and reverse primers (primer
sequences are shown in Table 1). An initial denaturation step at 95
oC for 10 min was followed by 40 cycles of amplification alternating between
95 oC for 30 s, the corresponding annealing
temperature for each gene for 30 s and 72 oC
for 30 s. Each sample was assayed in triplicate. The relative amount of target transcripts
was normalized to the number of human HPRT transcripts found in the same sample.
They were quantified using an efficiency corrected quantification model to obtain
the derivative ratio values:
| Gene | Primer (5'-3') | Tª 1 (°C) | Bp2 | |
| Forward | Reverse | |||
| HPRT | ATGGTGGGTTGTGCTTTCC | AGTTGTGTTTGTGCTTTCTGCT | 60 | 123 |
| Gli1 | CAGTGTGGGGACAGAAGGA | CGGGGAGAAGAAAAGAGTGG | 57.4 | 132 |
| Gli3 | ATGGACCCCAGGAATGGT | CAACCTTCTTGCTCACACATGTAAG | 64 | 167 |
| Ptch1 | CTTCGCTCTGGAGCAGATTT | CAGGACATTAGCACCTTCT | 55 | 354 |
| Cyclin D2 | GAAGGACATCCAACCCTACA | TCTTCTTCGCACTTCTGTTCC | 58.5 | 81 |
| Plakoglobin | ACGCTCAAGAAAACCACCAC | GTCCTCGCCTGACACACC | 58 | 132 |
| Nkx2.2 | TCTACGACAGCAGCGACAAC | CGCCTTGGAGAAAAGCACTC | 60 | 221 |
| Smo | AAGGCTGCACGAATGAGGT | GGGTTCTGGCACTGGATG | 57 | 133 |
| Sufu | CCTCCAGATCGTTGGTGTCT | CCCCTCCGCATGTCAGTT | 65 | 128 |
| Pax6 | ATGCCTCATAAAGGGGGAAG | AGTTGTGTTTGTGCTTTCTGCT | 60 | 112 |
| Bmi-1 | ATGCCTCATAAAGGGGGAAG | AGTTGTGTTTGTGCTTTCTGCT | 60 | 195 |
| Notch1 | ATTGTTCGTTACCTGGAGACC | GGCAGCATCAGCAGAAGG | 64 | 243 |
| Notch2 | GAGGCGACCGAGAAGATG | CCTGTGCCATTGTGGTAGG | 63.7 | 338 |
| Myc-N | CCCTGAGCGATTCAGATGAT | AATGTGGTGACAGCCTTGGT | 64.3 | 158 |
1Annealing temperatures.
2 Base pairs of the amplified product of primers.
MTT assay and cyclopamine treatment
MTT assay was performed as follows. After preparing serial dilutions of cells in
culture medium 5 × 103 cells were plated
out, in triplicate, in 200 µl of the dilutions (medium and cyclopamine in DMSO)
into wells of a micro-titer plate. Eight control wells of medium alone were included
to provide the blanks for absorbance readings. Cells were incubated under growing
conditions with 15 µM cyclopamine (Sigma Aldrich, UK) dissolved in DMSO for 24 and
48 h. Fifty µl of 3-(4, 5-dimethylthiazol-2)-2, 5-diphenyltetrazolium bromide (MTT)
reagent were added to each well, including controls. Then, plates were further incubated
for 2–4 h. When the purple precipitate was clearly visible under the microscope,
we removed the medium from each well and added 100 µl of detergent reagent (DMSO)
to all wells, including controls. After gently swirling (not shaking), we left plates
for 15-30 min at room temperature. The absorbance was measured in each well, including
the blanks, at 570 nm in a micro-titer plate reader (Perkin-Elmer, Boston, MA).
The experiment was performed for three independent times. In our experiment model
for cyclopamine treatment, cells cultured in normal medium were seeded in 6-well
plates at a density of 250,000 cells/well. After 24 h, the medium was changed to
low serum condition (2.5% FBS). Then, the cells were treated with 15 µM of cyclopamine
dissolved in DMSO for 48 h. Cells were also treated with DMSO alone as control.
The amount of DMSO never exceeded the 1% of the total volume as 1% DMSO alone showed
no cytotoxicity.
Gli1 siRNA knockout
For the transfection study, we took the Daoy medulloblastoma cell line grown in
supplemented RPMI media with 2% fetal bovine serum at 37
oC with 5% CO2. Moreover, we obtained
a pool of siRNAs against Gli1 gene from Stealth™ (Invitrogen, Carlsbad, CA) and
mixed them: Gli1 siRNA:
GCACAUACCUGCUUCGGGCAAGAUAU (GLI-HSS104170),
AUAUCUUGCCCGAAGCAGGUAGUGC (GLI-HSS104170),
CAGCUUGGACGAGGGACCUUGCAUU (GLI-HSS178441),
AAUGCAAGGUCCCUCGUCCAAGCUG (GLI-HSS178441),
GGGAUGGCUGCAGCCAGGAAUUUGA (GLI-HSS178442),
UCAAAUUCCUGGCUGCAGCCAUCCC (GLI-HSS178442).
We delivered siRNAs against Gli1 (100 nM/well) and its negative control to the Daoy cell line using Lipofectamine™ 2000 as a transfection reagent and Opti-MEM® I reduced serum media. The transfections were conducted on six well plates. RNA was extracted from transfected, negatively transfected and untransfected cells after 48 h. The extracted RNA was used for further analysis of expression levels of the downstream target genes of the Shh pathway. The experiment was performed in triplicate.
Scratching assay for cell migration
The Daoy monolayers from untransfected or untreated, DMSO or Lipofectamine 2000
and cyclopamine or siRNA-treated dishes were suspended by treatment with trypsin/EDTA
(Gibco) and approximately 2.5 × 105 cells
in 2 ml of medium were seeded into wells of a 6-well plate and grown for 24 h to
about 90% confluent in medium containing 10% FBS. Next, cell migration was assayed
by evaluating closure of a linear wound produced by scratching the cell monolayer
on a 6-well plate in 2.5% fetal bovine serum plus medium. The open gap was then
inspected microscopically over time as the cells move in and fill the damaged area.
To analyse the cell migration we took pictures at 0, 6, 12, 24 and 48 h and the
cells were kept in the incubator at 37 oC
in an atmosphere with 5% CO2 while waiting
between pictures. Therefore, we evaluated cell migration in pre-treated cells.
Colony formation assay in culture
This assay was performed to evaluate the ability of pre-treated Daoy cells (at the
same conditions as before) to create a colony in monolayer cultures. Given that
Daoy cells grow adhered to a layer, the test would take less time than in soft agar.
2.5 × 102 Daoy cells per well of both conditions
were cultured in a 6-well plate for 10 days. After that, we discarded the medium,
cells were fixed 30 minutes with 4% paraformaldehyde. They were then stained with
1% Crystal Violet (Sigma Aldrich, UK) for 15 min and counted. The experiment was
performed in triplicate.
Colony formation assay in soft agar
Pre-treated Daoy cells were mixed with 0.4% Noble agar (in Dulbecco’s modified Eagle’s
medium supplemented with 10% fetal bovine serum) and plated at 5000 cells/well into
6-well plates containing a solidified bottom layer (1% Noble agar in the same growth
medium). After 20 days, colonies were stained with 250 µL of 2.5% Crystal Violet
(Sigma Aldrich, UK) for 5 min, washed 4-5 times with PBS to remove excess of dye
and finally photographed. The experiment was performed for 3 independent times for
the cell line cultured in medium with or without cyclopamine in DMSO or siRNA in
Lipofectamine 2000.
Statistical analysis
Statistical analysis was performed using SPSS Statistical Software (Chicago, IL,
USA). The data represent mean ± SEM. One-way analysis of variance (ANOVA) was used
to compare the controls and treated or transfected group of three independent soft
agar clonogenic assays. The same analysis was performed in MTT assay comparing cyclopamine
treatment with untreated cells. Differences were considered statistically significant
when the p value was less than 0.05 and highly significant when the p value was
less than 0.01. Remaining studies were performed in triplicate.
ResultsTop
Cyclopamine and Gli1 siRNA reduce Daoy cell viabilityWe investigated the concentration of cyclopamine that could inhibit cell growth in these cells. The MTT cell proliferation assay was performed to measure the cell proliferation rate when Daoy cells were treated with cyclopamine at different concentrations (from 1 to 50 µM). This assay allowed us to obtain the IC50 of cyclopamine to be used for next cellular and molecular studies. We took 15 µM as cyclopamine IC50 as it was the closest and less concentrated value to the real IC50 (Figure 2). In addition, this concentration showed the same effect at 24 or 48 h. The differences observed from 10 to 50 µM were highly significant. The amount of DMSO never exceeded 1% of the total reaction volume.
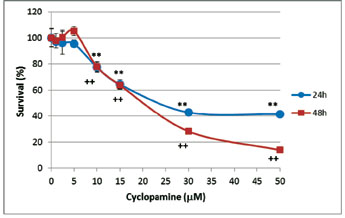
Figure 2 Cyclopamine inhibits growth of Daoy cell line. Sensitivity of Daoy cells to different concentrations of cyclopamine (1; 2.5; 5; 10; 15; 30 and 50 µM) dissolved in DMSO compared to controls (none or DMSO-treated cells) was assessed by the MTT assay. Data reflect the means ± SEM of three independent experiments. **P < 0.01 for 24 h and ++P < 0.01 for 48h.
Downregulation of Gli1 expression in Daoy by siRNA or cyclopamine
We evaluated the effects of direct silencing of Gli1 expression on cell behavior
by transfecting cells with a three-pool Gli1 siRNA or a control siRNA for 48 h (Figure
3A). In addition, we compared Gli1 siRNA with a direct inhibitor of Smo receptor
(cyclopamine) that acts upstream of Gli1 [23]. Surprisingly, the chemical approach,
15 µM of cyclopamine, resulted in 87% reduction of Gli1 mRNA expression in Daoy
cells compared to untreated cells and around 70% reduction compared to DMSO (Figure
3B). Gli1 siRNA decreased 94% of Gli1 mRNA levels (Figure 3A).
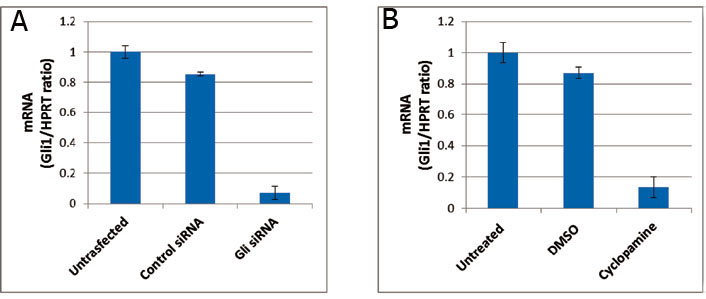
Figure 3 Knockdown of Gli1. (A) Gli1 siRNA highly reduces Gli1 in Daoy cell line. (B) Cyclopamine inhibition produces a Gli1 expression which is two-fold Gli1 mRNA levels managed by Gli1 siRNA as determined by quantitative RT-PCR (representative of triplicate experiments). Values represent the mean levels of Gli1 mRNA ± SEM of untransfected/untreated and Gli1 siRNA/cyclopamine in DMSO-treated cells related to control siRNA/DMSO (cyclopamine solvent)-treated Daoy cells. The derivative ratio values describe the relative expression change of the target gene relative to the HPRT reference gene expression.
Shh inhibition reduces in vitro tumorigenesis of Daoy cells
We cultured non-transfected and siRNA transfected Daoy cells, either control or
Gli1, in 6-well plates where they anchored. After 10 days, we detected less number
of colonies (Figure 4A) that corresponded to a 70% decrease in colony formation
(Figure 4B, left panel). Likewise, cells treated with cyclopamine or with DMSO showed
similar results (Figure 4B, right panel).
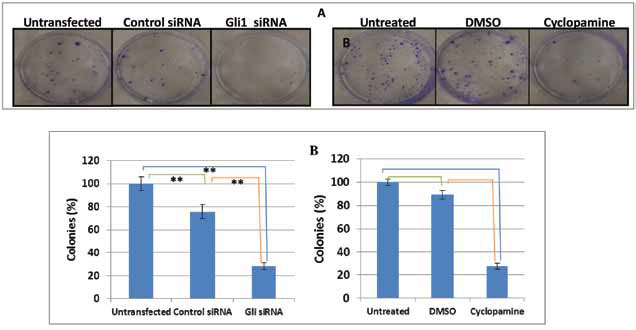
Figure 4 Effects of Gli1 downregulation on the ability to form colonies. (A) The colonies formed after 10 days were stained with 0.5% crystal violet solution (left and right panels). (B) Number of grown colonies of all conditions was counted visually (left and right panels). Data reflect the means ± SEM of triplicate experiments (*p < 0.05; **p < 0.01).
In the same way, we sought to determine the tumor formation ability using an in vitro 3-dimensional colony formation assay in soft agar media (Figure 5) to better simulate the process of in vivo carcinogenesis.
This analysis demonstrated that cells underexpressing Gli1 (by either siRNA or cyclopamine) formed distinct microscopic three-dimensional colonies embedded in different layers of soft agar (Figure 5A). The number of colonies was counted under a microscope and plotted in Figure 4B. Interestingly, there were very few small colonies in the plates containing untransfected or untreated cells as well as control siRNA and DMSO treatment and almost no colonies were observed in the Gli1 siRNA transfected or cyclopamine treated Daoy cells (Figure 5A-B). These observations demonstrated that Gli1 downregulation reduced in vitro tumorigenesis of Daoy cells. The differences found between groups were highly significant. The reduction in number of colonies and sizes of the 3D-growth demonstrated potential roles of Gli1 in 3D-colony formation and tumorigenesis.
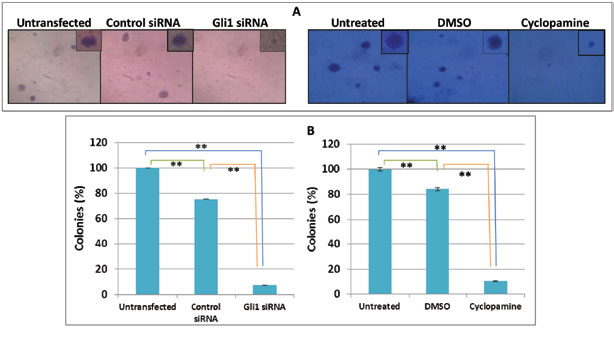
Figure 5 Cyclopamine and siRNA highly prevent tumorigenic growth. (A) Microscopic view of colonies with a magnification of x100 (big images) and x250 (their expanded pictures after 20 days). (B) Percentages of colonies were obtained after counting five-microscopic fields of each condition and compared with untransfected/treated control. Data reflect the means ± SEM of triplicate experiments. **p < 0.01.
Gli1 silencing impairs Daoy cell migration
Since targeting Gli1 inhibited cell viability and decreased tumorigenic growth in
Daoy cells, we investigated the effect of Gli1 silencing on the migration potential
of Daoy cells using an in vitro wound-healing assay, frequently used as a simple
assay to mimic and assess migration [24] . In this assay, an artificial wound is
created by scratching a confluent monolayer of cells, while the ability of the cells
to move into and close the wound is thought to predict their migration ability in
vivo [24] . Therefore, after culturing Daoy cells from all six conditions (untransfected;
control siRNA; Gli1 siRNA and untreated; DMSO; cyclopamine in DMSO) for 24 h they
were scratched. From that moment (0 h) we evaluated Daoy migration ability for 48
h. Here, we can see a huge different behavior between controls and siRNA or cyclopamine
treatments supporting our hypothesis (Figure 6). Surprisingly, cyclopamine did not
only decrease cell migration, but also diminished initial confluence (Figure 6,
right panel).
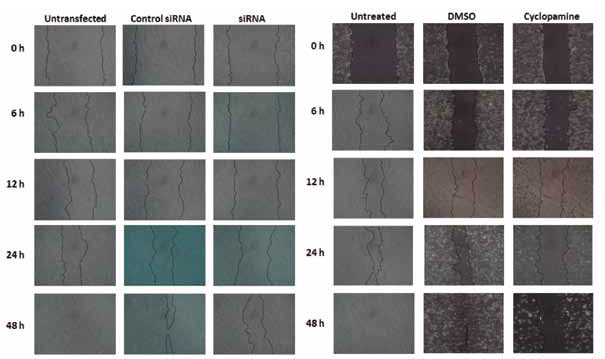

Figure 6 Scratch wound assay for cell migration determination. Both, 100 nM of Gli1 siRNA (left panel) and 15 µM cyclopamine (right panel) decrease cell migration. Cells were untransfected or transfected by control siRNA and Gli1 siRNA (left panel), while right panel shows untreated and DMSO (vehicle) or cyclopamine in DMSO-treated Daoy cells. Similar results were obtained in triplicate.
qRT-PCR expression of Shh pathway components and related genes
After transfection with a three-pool siRNA against Gli1 and treatment with cyclopamine,
we first validated the success on downregulating Gli1 by qRT-PCR. Next, we attempted
to determine the expression of Gli1 target genes, including Ptch1, Cyclin D2, Plakoglobin,
Pax6, Nkx2.2 and Bmi-1. Furthermore, we analysed components of Shh signaling pathway
such as Smo, Sufu and Gli3. Focusing on Shh pathway crosstalks that have been reported
recently to be related with N-myc, Notch1 and Notch2 [25, 26]. Here, we found that
Gli1 inhibition by siRNA or Smo antagonist produced downregulation of Ptch1, Cyclin
D2, Plakoglobin, Nkx2.2, Bmi-1, Smo and N-myc (Figure 7A, B, C, D, E, G and J respectively).
With connection with Sufu and Gli3, they showed parallel results, where Gli1 siRNA
did neither decrease nor increase expression of both genes, whereas cyclopamine
reduced them in 15-25% (Figure 7H and I). Pax6 and Notch1 mRNA levels were up-regulated
by either Gli1 siRNA or cyclopamine (Figure 7F and 7K). However, Notch2 was decreased
by Gli1 siRNA and increased by cyclopamine (Figure 7L). Percentage data on expression
changes after Gli1 knock-down appear on Table 2.
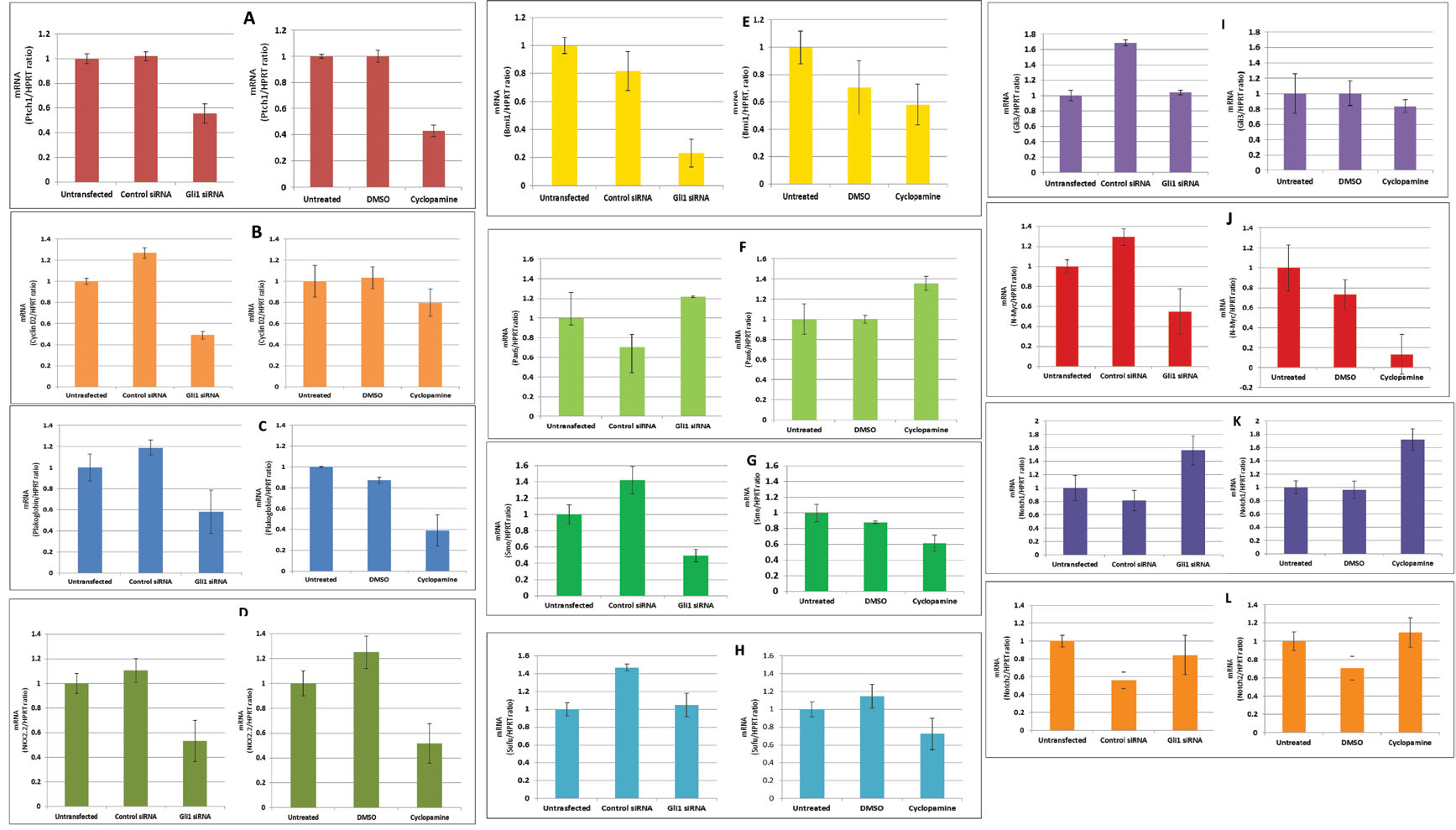
Figure 7 qRT-PCR of Ptch1, Cyclin D2, Pax6, Plakoglobin, Nkx2.2, Bmi-1, Smo, Sufu, Gli3, N-myc, Notch1 and Notch2 genes in Gli1 knockdown in Daoy MB cell line. The derivative ratio values describe the relative expression change of the target gene relative to the HPRT reference gene expression. Downregulation of Ptch1, Cyclin D2, Plakoglobin, Nkx2.2, Bmi1, Smo and N-myc was detected after Shh inhibition. Sufu and Gli3 showed parallel results, where Gli1 siRNA did neither decrease nor increase expression of both genes, whereas cyclopamine reduced them in 15-25%. Pax6 mRNA levels were upregulated by either Gli1 siRNA or cyclopamine. Finally, Notch1 was upregulated after inhibition of SHH, while Notch2 showed contrasting results, as siRNA inhibition decreased its expression, while cyclopamine induced upregulation of Notch2 (See Table 2 for quantification of expression results).
| Gene | GLI1 knock-down after siRNA treatment or cyclopamine1 (%) | |
Gli1 siRNA |
Cyclopamine |
|
| Ptch1 | - 43 |
- 58 |
| Cyclin D2 | - 50 |
- 20 |
| Plakoglobin | - 40 |
- 60 |
| Nkx2.2 | - 50 |
- 50 |
| Bmi-1 | - 75 |
- 40 |
| Pax6 | + 20 |
+ 40 |
| Smo | - 50 |
- 60 |
| Sufu | + 5 |
- 25 |
| Gli3 | + 5 |
- 15 |
| Myc-N | - 45 |
- 85 |
| Notch1 | + 56 |
+ 65 |
| Notch2 | - 18 |
+ 10 |
1(+) Increase in the expression of each
gene after Gli1 siRNA or cyclopamine treatment.
(-) Decrease in expression respect to controls.
DiscussionTop
Sonic hedgehog signals through a cascade of downstream effectors, most of which are still unknown. Yet, to our knowledge, there are no highly effective approved targeted therapies to treat Shh-driven medulloblastoma. However, vismodegib, an oral inhibitor of the hedgehog pathway, is being studied in an ongoing clinical trial in adult patients with recurrent, progressive, or refractory to standard therapy medulloblastoma [27] . In addition, although it is unclear what side effects may occur in children, as permanent defects in bone growth have been seen in mouse models of Hh inhibitors [28] , few clinical trials have been started in this population with recurrent or refractory medulloblastoma [29, 30]. Moreover, vismodegib became on January 30, 2012 the first drug approved by the US Food and Drug Administration (FDA) for the treatment of adults with metastatic basal cell carcinoma, or with locally advanced basal cell carcinoma that has recurred following surgery or who are not candidates for surgery and who are not candidates for radiation [31] . This situation may stem from our limited knowledge of which proteins are directly responsible for mediating Shh signaling in this tumor type and, hence, serve as effective therapeutic targets. Individuals with somatic mutations of Ptch, Smo, and Sufu, as well as amplifications of Gli1 have been found to be predisposed to sporadic medulloblastoma [32] . Moreover, it is reported that 1/200 spontaneous abortions are due to mutation in Shh and Ptch. As illustrated in this study, the role of Gli1 transcription factor is significant for tumorigenicity in Daoy cells. Hence, inhibition of Gli1 activity within a tolerated therapeutic window may provide a successful pharmacological approach to treat Shh-positive medulloblastomas. In this perspective, we report a role for the transcription factor Gli1 in the growth, survival, migration and expression of related signaling in Daoy medulloblastoma cell line.
Here, we evaluated cyclopamine inhibition as an approach to inhibit the Gli1 trigger (Smo) and compared with direct Gli1 downregulation by a three-pool siRNA. An initial study demonstrated that cyclopamine decreased Daoy cell viability in a dose-dependent manner in vitro. In this assay we found that IC50 of cyclopamine for Daoy cells was around 15 µM and we took that concentration as the closest one tested. Similar results were chosen in a study performed with PNET cell lines, including Daoy [35] .
However, response to cyclopamine varied among pancreatic adenocarcinoma cell lines with IC50 values differing over 5-fold (from ˜9 to 45 µM) [36] . Kandala and Srivastava [37] reported the treatment of ovarian cancer cells with 25 µM cyclopamine, although they also used 10 µM. These differences may be due to differences in format and time assessed as one report used cyclopamine under suspension culture conditions [37] .
The data from Gli1 siRNA silencing and Smo antagonizing studies suggest that the chemical approach is adequate to target Shh pathway. Notwithstanding, it was twofold increased comparing with the three-pool siRNA against Gli1 used. Other study which focused on a direct Gli1 inhibitor, GANT58, which has been reported to block Gli1-induced transcription in vitro showed lower capacity to diminish Gli1 expression than cyclopamine [38] . Taken together, our findings might indicate that the Shh-Gli1 pathway remains active in Daoy cell line by either a mutation in Ptch or Smo, or eventually by high expression of Shh as an autocrine signaling, and not by Sufu dysregulation.
Besides, we show that downregulation of Gli1, by any of both methods, greatly diminishes colony forming ability in adherent and nonadherent conditions in terms of both quantity and colony size in the medulloblastoma cell line. In particular, we observed a decrease of 75% in number of colonies by Gli1 siRNA compared with untransfected cells when cells were cultured attached, and even a reduction of 15% more in unattached assays using soft agar. Similar results were obtained with cyclopamine treatment. A previous study found cyclopamine more effective in prostate carcinoma xenograft models than in prostate carcinoma cell line 22Rv1 in vitro [33] . In consistence with that, we suggest that due to the greater difference in soft agar than in the attached assay and that independent growth assay in soft agar is assumed to be closely related to the process of in vivo carcinogenesis, cyclopamine might show better results in tumor growth in a xenograft assay with regard to the surrounding stroma.
Medulloblastoma has been described as an invasive and a rapidly growing tumor that, unlike most brain tumors, spreads through the cerebrospinal fluid (CSF) and frequently metastasizes to different locations in the brain and spine. Further, we observed that the migration ability of the Daoy cells when Gli1 was silenced was dramatically reduced compared with untransfected or untreated cells as assessed in conventional scratch assays. Moreover, we observed that cells treated with cyclopamine not only diminished migration, but also decreased initial confluence. Although this could be mediated also by the effect of DMSO, solvent of cyclopamine, a huge difference can be detected intra-experimentally (between DMSO and cyclopamine) and inter-experimentally (between Gli1 siRNA and cyclopamine).
In the course of this study, we sought to understand the regulation of certain downstream target genes and other genes related with the Shh pathway in the Daoy cell line. In addition, we attempted to explore any putative regulation of these genes by the major transcription factor involved in Shh signaling, Gli1. Among the Gli1 target genes studied we found that Ptch1, Cyclin D2, Plakoglobin, Nkx2.2, Bmi-1 were downregulated after both treatments. Especially, Ptch1 and Plakoglobin mRNA levels showed less expression using cyclopamine. This result does not corroborate a previous report which suggests that Gli1 downregulates Plakoglobin in Gli1 transformed epithelial cells [39] . For Cyclin D2, our finding is supported by the previous report showing similar results in Gli1 transformed epithelial cells [39]. From Nkx2.2 result, we support the conclusion of a report [40] which suggests that Gli1 up-regulates Nkx2.2. Bmi-1 is a stem cell marker gene linked to Shh [41] that is up-regulated as Nkx2.2 in medulloblastoma. But treatment of the Shh pathway makes Nkx2.2 and Bmi-1 mRNA levels diminish. Nevertheless Pax6, another target gene, presented higher mRNA level than untransfected or untreated controls in agreement with this work [42] involving the role of Gli1 in downregulating its expression. Focusing on Smo, qRT-PCR revealed an mRNA reduction between 40-50% by both approaches. Gli3 and Sufu tumor suppressors showed opposite mRNA values. While inhibition by Gli1 siRNA produced a subtle increase, the inhibition by the Smo antagonist resulted in a decrease of Gli3 and Sufu mRNA levels, result that could be explained by the difference on Gli1 mRNA levels obtained.
Finally, in relation with the Shh pathway and other cancer pathways, N-myc and Notch, we found encouraging results related with prior works. It has been found that continued N-myc expression in granule neuron progenitors is a key factor contributing to Shh-driven medulloblastoma tumorigenesis [43] . N-myc was downregulated by Gli1 or Smo targeting and they could potentially affect stem-like cells in the tumors [44] . Here, we observed that cyclopamine inhibition produced twofold N-myc mRNA levels managed by Gli1 siRNA. This makes us think that Smo plays, in normal conditions, activating N-myc expression before Gli1. In connection with Notch1 and 2, it is known that Notch1 is associated with cell cycle exit and differentiation, whereas Notch2 promotes proliferation of cerebellar granule neuron progenitors [45] . Parallel results were found in Notch1 expression when targeting Smo or Gli1, obtaining an increase in Notch1 mRNA level. In particular, we found that Notch2 showed opposite outcomes than Notch1. Gli1 siRNA produced a decrease of Notch2 mRNA unlike cyclopamine which increased Notch2 expression. Hence, we suggest that this difference in Notch2 mRNA levels can be explained by the fact that cyclopamine inhibition produced twofold Gli1 mRNA levels managed by Gli1 siRNA.
To sum up, this work represents a quick overview of the Shh pathway’s role and its connections with other signals related with carcinogenesis in medulloblastoma. Two ways to inhibit Shh were tested. Inhibition led to in vitro tumorigenesis decrease in terms of cell viability, migration and colony formation in culture and in soft agar. Further studies remain to be done for a better understanding of the crosstalks to provide more relevant information toward the development of effective targeted therapies for the treatment of Shh-driven medulloblastoma.
Funding
This research was supported in part by grants from the Departmento de Salud del Gobierno de Navarra, Caja Navarra (project 13912), Fundación Universitaria de Navarra, Pamplona; and Fondo de Investigación Sanitaria (PI081849, to JSC; and PI-101972, to JAR), Madrid.
Conflict of interest
The authors wish to express that they have no conflict of interest.
Acknowledgements
We thank Pablo Sáez for his help and advice with the artwork.
ReferencesTop
[1] Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, et al. (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114: 97–109. Article Pubmed
[2] Marino S (2005) Medulloblastoma: developmental mechanisms out of control. Trends Mol Med 11:17–22. Article Pubmed
[3] Gatta G, Zigon G, Capocaccia R, Coebergh JW, Desandes E, et al. (2009) Survival of European children and young adults with cancer diagnosed 1995–2002. Eur J Cancer 45:992–1005. Article Pubmed
[4] Taylor MD, Northcott PA, Korshunov A, Remke M, Cho YJ, et al. (2012) Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol 123:465–472. Article Pubmed
[5] Thompson MC, Fuller C, Hogg TL, Dalton J, Finkelstein D, et al. (2006) Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J Clin Oncol 24:1924–1931.Article Pubmed
[6] Massimino M, Giangaspero F, Garr M, Gandola L, Poggi G, et al. (2011) Childhood medulloblastoma. Crit Rev Oncol 79: 65–83. Article Pubmed
[7] Zurawel RH, Chiappa SA, Allen C, Raffel C (1998) Sporadic medulloblastomas contain oncogenic beta-catenin mutations. Cancer Res 58:896–899. Article Pubmed
[8] Eberhart CG, Tihan T, Burger PC (2000) Nuclear localization and mutation of beta-catenin in medulloblastomas. J Neuropathol Exp Neurol 59:333–337. Article Pubmed
[9] Cho YJ, Tsherniak A, Tamayo P, Santagata S, Ligon A, et al. (2011) Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J Clin Oncol 29:1424–1430. Article Pubmed
[10] Kool M, Korshunov A, Remke M, Jones DT, Schlanstein M, et al. (2012) Molecular subgroups of medulloblastoma: an international meta-analysis of transcriptome, genetic aberrations, and clinical data of WNT, SHH, Group 3, and Group 4 medulloblastomas. Acta Neuropathol 123:473–484. Article Pubmed
[11] Wechsler-Reya R, Scott MP (2001) The developmental biology of brain tumors. Annu Rev Neurosci 24:385–428. Article Pubmed
[12] Hatten ME (1999) Expansion of CNS precursor pools: a new role for Sonic Hedgehog. Neuron 22:2–3. Article Pubmed
[13] Kenney AM, Cole MD, Rowitch DH (2003) Nmyc upregulation by sonic hedgehog signaling promotes proliferation in developing cerebellar granule neuron precursors. Development 130:15–28. Article Pubmed
[14] Yoon JW, Kita Y, Frank DJ, Majewski RR, Konicek BA, et al. (2002 ) Gene expression profiling leads to identification of Gli1-binding elements in target genes and a role for multiple downstream pathways in Gli1-induced cell transformation. J Biol Chem 277: 5548–5555. Article Pubmed
[15] Pietsch T, Waha A, Koch A, Kraus J, Albrecht S, et al. (1997) Medulloblastomas of the desmoplastic variant carry mutations of the human homologue of Drosophila patched. Cancer Res 57:2085–2088. Article Pubmed
[16] Lee Y, Kawagoe R, Sasai K, Li Y, Russell HR, et al. (2007) Loss of suppressor-of-fused function promotes tumorigenesis. Oncogene 26:6442–6447. Article Pubmed
[17] Hallahan AR, Pritchard JI, Hansen S, Benson M, Stoeck J, et al. (2004) The SmoA1 mouse model reveals that notch signaling is critical for the growth and survival of sonic hedgehog-induced medulloblastomas. Cancer Res 64:7794–7800. Article Pubmed
[18] Briscoe J, Sussel L, Serup P, Hartigan-O’Connor D, Jessell TM, et al. (1999) Homeobox gene Nkx2.2 and specification of neuronal identity by graded Sonic hedgehog signalling. Nature 398:622–627. Article Pubmed
[19] Ingram WJ, Wicking CA, Grimmond SM, Forrest AR, Wainwright BJ (2002 ) Novel genes regulated by Sonic Hedgehog in pluripotent mesenchymal cells. Oncogene 21: 8196–8205. Article Pubmed
[20] Leung C, Lingbeek M, Shakhova O, Liu J, Tanger E, et al. (2004) Bmi-1 is essential for cerebellar development and is overexpressed in human medulloblastomas. Nature 428:337–341. Article Pubmed
[21] Chuang PT, McMahon AP (1999) Vertebrate Hedgehog signalling modulated by induction of a Hedgehog-binding protein. Nature 397:617–621. Article Pubmed
[22] Zurawel RH, Allen C, Chiappa S, Cato W, Biegel J, et al. (2000) Analysis of PTCH/SMO/SHH pathway genes in medulloblastoma. Genes Chromosomes Cancer 27: 44–51. Article Pubmed
[23] Chen JK, Taipale J, Cooper MK, Beachy PA (2002) Inhibition of Hedgehog signaling by direct binding of cyclopamine to Smoothened. Genes Dev 16:2743–2748. Article Pubmed
[24] Liang CC, Park AY, Guan JL (2007) In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc 2:329–333. Article Pubmed
[25] Thomas WD, Chen J, Gao YR, Cheung B, Koach J, et al. (2009) Patched1 deletion increases N-Myc protein stability as a mechanism of medulloblastoma initiation and progression. Oncogene 28:1605–1615. Article Pubmed
[26] Sivakumar KC, Dhanesh SB, Shobana S, James J, Mundayoor S (2011) A systems biology approach to model neural stem cell regulation by notch, shh, wnt, and EGF signaling pathways. OMICS 15:729–737. Article Pubmed
[27] U.S. National Institutes of Health (2012) Study of vismodegib in combination with temozolomide versus temozolomide alone in patients with medulloblastomas with an activation of the Sonic Hedgehog Pathway. http://clinicaltrials.gov/ct2/show/study/NCT01601184?term=vismodegib&rank=5
[28] Kimura H, Ng JM, Curran T (2008) Transient inhibition of the Hedgehog pathway in young mice causes permanent defects in bone structure. Cancer Cell 13:249–260. Article Pubmed
[29] U.S. National Institutes of Health (2012) GDC-0449 in treating young patients with medulloblastoma that is recurrent or did not respond to previous treatment. http://clinicaltrials.gov/ct2/show/study/NCT00822458?term=vismodegib&age=0&phase=0&rank=1
[30] U.S. National Institutes of Health (2012) GDC-0449 in treating younger patients with recurrent or refractory medulloblastoma. http://clinicaltrials.gov/ct2/show/study/NCT01239316?term=vismodegib&rank=18
[31] U.S. Food and Drug Administration (2012) FDA approves new treatment for most common type of skin cancer. http://www.fda.gov/NewsEvents/Newsroom/PressAnnouncements/ucm289545.htm
[32] Northcott PA, Hielscher T, Dubuc A, Mack S, Shih D, et al. (2011) Pediatric and adult sonic hedgehog medulloblastomas are clinically and molecularly distinct. Acta Neuropathol 122:231–240. Article Pubmed
[33] Northcott PA, Nakahara Y, Wu X, Feuk L, Ellison DW, et al. (2009) Multiple recurrent genetic events converge on control of histone lysine methylation in medulloblastoma. Nat Genet 41:465–472. Article Pubmed
[34] Taylor MD, Liu L, Raffel C, Hui CC, Mainprize TG, et al. (2002) Mutations in SUFU predispose to medulloblastoma. Nat Genet 31:306–310. Article Pubmed
[35] Enguita-Germán M, Schiapparelli P, Rey JA, Castresana JS (2010) CD133 cells from medulloblastoma and PNET cell lines are more resistant to cyclopamine inhibition of the sonic hedgehog signaling pathway than CD133- cells. Tumour Biol 31:381–390. Article Pubmed
[36] Steg A, Amm HM, Novak Z, Frost AR, Johnson MR (2010) Gli3 mediates cell survival and sensitivity to cyclopamine in pancreatic cancer. Cancer Biol Ther 10:893–902. Article Pubmed
[37] Kandala PK, Srivastava SK (2012) Diindolylmethane-mediated Gli1 protein suppression induces anoikis in ovarian cancer cells in vitro and blocks tumor formation ability in vivo. J Biol Chem 287:28745–28754. Article Pubmed
[38] Lauth M, Bergström A, Shimokawa T, Toftgård R (2007) Inhibition of GLI-mediated transcription and tumor cell growth by small-molecule antagonists. Proc Natl Acad Sci U S A 104:8455–8460. Article Pubmed
[39] Yoon JW, Kita Y, Frank DJ, Majewski RR, Konicek BA, et al. (2002) Gene expression profiling leads to identification of Gli1-binding elements in target genes and a role for multiple downstream pathways in Gli1-induced cell transformation. J Biol Chem 277: 5548–5555. Article Pubmed
[40] Fuccillo M, Joyner AL, Fishell G (2006) Morphogen to mitogen: the multiple roles of hedgehog signalling in vertebrate neural development. Nat Rev Neurosci 7:772–783. Article Pubmed
[41] Leung C, Lingbeek M, Shakhova O, Liu J, Tanger E, et al. (2004) Bmi-1 is essential for cerebellar development and is overexpressed in human medulloblastomas. Nature. 428:337–341. Article Pubmed
[42] Ericson J, Rashbass P, Schedl A, Brenner-Morton S, Kawakami A, et al. (1997) Pax6 controls progenitor cell identity and neuronal fate in response to graded Shh signaling. Cell 90:169–180. Article Pubmed
[43] Solecki DJ, Liu XL, Tomoda T, Fang Y, Hatten ME (2001) Activated Notch2 signaling inhibits differentiation of cerebellar granule neuron precursors by maintaining proliferation. Neuron 31: 557–568. Article Pubmed
[44] von Bueren AO, Shalaby T, Rajtarova J, Stearns D, Eberhart CG, et al. (2007) Anti-proliferative activity of the quassinoid NBT-272 in childhood medulloblastoma cells. BMC Cancer 7:19. Article Pubmed
[45] Solecki DJ, Liu XL, Tomoda T, Fang Y, Hatten ME (2001) Activated Notch2 signaling inhibits differentiation of cerebellar granule neuron precursors by maintaining proliferation. Neuron 31: 557–568. Article Pubmed


