Journal of Cancer Research & Therapy
An International Peer-Reviewed Open Access Journal
ISSN 2052-4994


- Download PDF
- |
- Download Citation
- |
- Email a Colleague
- |
- Share:
-
- Tweet
-

Journal of Cancer Research & Therapy
Volume 8, Issue 2, April 2020, Pages 9–16
Original researchOpen Access
MiR-381-3p inhibits proliferation, invasion and migration of hepatocellular carcinoma
-
Wu Changtao1, Mingxin Liu2, Du Xiufang3, Zhao Yuan4, Zhao Chang5, Chen Jie4, Liu Junjie6, Tang Weizhong7,*
 and
Zhang Chunyan8,*
and
Zhang Chunyan8,*
- 1 Department of Colorectal and Anal Surgery, The First Affiliated Hospital, Guangxi Medical University, Nanning 530021, China
- 2 School of medicine, University of Electronic Science and Technology of China, Chengdu, Sichuan, China
- 3 Department of Pathology, The First Affiliated Hospital, Guangxi Medical University, Nanning 530021, China
- 4 Department of Hepatobiliary Surgery, Affiliated Tumor Hospital of Guangxi Medical University, Nanning 530021, China
- 5 Department of Interventional Radiology, Affiliated Tumor Hospital of Guangxi Medical University, Nanning 530021, China
- 6 Department of Ultrasound, Affiliated Tumor Hospital of Guangxi Medical University, Nanning 530021, China
- 7 Department of Gastrointestinal Surgery, Affiliated Tumor Hospital of Guangxi Medical University, Nanning 530021, China
- 8 Department of Experimental Research, Affiliated Tumor Hospital of Guangxi Medical University, Nanning 530021, China
*Corresponding authors: Tang Weizhong, Department of Gastrointestinal Surgery, Affiliated Tumor Hospital of Guangxi Medical University, Hedi Road, Nanning 530021, People’s Republic of China, Tel: +86 0771 391 9631; Fax: +86 0771 391 9631; Email: tangweizhong@gxmu.edu.cn; Zhang Chunyan, Department of Experimental Research, Affiliated Tumor Hospital of Guangxi Medical University,71 Hedi Road, Nanning 530021, People’s Republic of China, Tel: +86 771 533 0855; Fax: +86 771 531 2000; Email: zhangchunyangmu@163.com
Received 12 December 2019 Revised 5 March 2020 Accepted 16 March 2020 Published 25 March 2020
DOI: http://dx.doi.org/10.14312/2052-4994.2020-2
Copyright: © 2020 Changtao W, et al. Published by NobleResearch Publishers. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
AbstractTop
MiR-381-3p is involved in the occurrence and development of various tumors. However, its biological roles in hepatocellular carcinoma (HCC) is still unknow. Here, we investigated the effects of miR-381-3p in HCC. qRT-PCR was used to detect the expression of miR-381-3p in HCC tissues and adjacent normal tissues. Results showed that miR-381-3p was down-regulated in 94 cases of HCC tissues. Clinical characteristics analysis showed that miR-381-3p expression was associated with gender, CA199, tumor size and metastasis of HCC patients. The expression level of miR-381-3p in SMMC-7721 HCC cells was regulated by transfection, and the effects of miR-381-3p on the function of HCC cells were detected by MTT proliferation assay, transwell assay and wound healing assay. And results showed that miR-381-3p inhibited the proliferation, invasion and migration of SMMC-7721 HCC cells. Multiple databases were used to predict the target genes of miR-381-3p, and GO enrichment analysis and KEGG pathway enrichment analysis were performed on these potential target genes by DAVID online analysis. The databases predicted 854 possible target genes of miR-381-3p, and analyzed their enrichment in three aspects of biological process, cellular component and molecular function. The strongest enrichment of KEGG pathway is the signaling pathways regulating pluripotency of stem cells. miR-381-3p inhibits HCC cell proliferation, invasion and migration and may be a new therapeutic target for HCC.
Keywords: hepatocellular carcinoma; miR-381-3p; proliferation; invasion; migration
IntroductionTop
Hepatocellular carcinoma (HCC) is one of the most common malignant tumors in the world, with a high mortality rate [1]. The number of HCC patients in China accounts for more than half of the global HCC cases [2]. HCC patients are not easily detected in the early stage, and most of them have already developed into advanced HCC when they are discovered. Liver resection (LR) and liver transplantation (LT) are the most commonly used treatment for early HCC, and more patients with advanced HCC have lost the best opportunity for surgical treatment, only non-surgical treatment such as transcatheter arterial chemoembolization and radioactive embolization can be selected [3, 4]. The recurrence rate of HCC is high, the overall survival time of HCC patients is low, and the prognosis is poor [5]. Therefore, looking for new methods for early diagnosis and treatment of HCC is crucial to improve the prognosis of patients.
MicroRNAs (miRNAs) are class of short non-coding RNAs with a length of about 22 nucleotides [6], which are involved in gene expression and play an important role in various processes of life activities [7]. Abnormal miRNA expression leads to a disorder of cell function, which may cause the occurrence of many diseases. At the same time, abnormal expression of miRNAs may also affect cancer-related key pathways, affecting various stages of tumorigenesis [8]. Ruan [9] et al. showed that miR-186 may be a new marker of lung cancer and inhibits lung cancer progression by targeting SIRT6. Wu [10] et al. found that miR-497 is low expression in many types of cancer, affecting the development of prostate cancer by down-regulating HDGF, and has a certain effect on the treatment of prostate cancer. The miRNA expression disorder also has an important impact on HCC [11]. Wong et al. found that miR-223 is low expressed in HCC patients and further affects the development of HCC by regulating STMN1, which may serve as a new potential target for the treatment of HCC [12].
In previous studies, miR-381-3p has been shown to play different roles in a variety of tumors. MiR-381-3p is down-regulated in tumor tissues such as bladder cancer, cervical cancer, and breast cancer, and affects the function of tumor cells. In this study, we aimed to explore whether miR-381-3p is also down-regulated in HCC tissues and whether it affects HCC cells function.
Materials and methodsTop
Tissue samples
The 94 HCC patients included in this study were from the Affiliated Tumor Hospital of Guangxi Medical University. None of the patients received radiotherapy or chemotherapy before surgery. The 94 HCC tissues and adjacent normal liver tissues were soaked in RNAlater (Qiagen, Valencia, CA, USA) and stored at -80o C. The study was approved by the Ethics Review Committee of the Affiliated Tumor Hospital of Guangxi Medical University. All patients provided informed consent and were interviewed by telephone or hospital until death or the last follow-up in May 2018. Their clinicopathologic features are listed in Table 1.
| Characters | miR-381-3p expression | Patients | OR(95%CI) | P value | ||
| Low (n=47) | High (n=47) | |||||
| Gender | 0.036 | |||||
| Female | 3 | 10 | 13 | Ref. 0.252(0.065-0.985) |
||
| Male | 44 | 37 | 81 | |||
| Age | 0.131 | |||||
| <60 | 34 | 40 | 74 | 0.458(0.164-1.277) | ||
| ≥60 | 13 | 7 | 20 | |||
| HBVDNA (cps/ml) | 0.636 | |||||
| <5.00*10e2 | 11 | 13 | 24 | 0.799(0.315-2.025) | ||
| ≥5.00*10e2 | 36 | 34 | 70 | |||
| AFP (ug/L) | 0.678 | |||||
| ≤400 | 22 | 20 | 42 | 1.188(0.526-2.681) | ||
| >400 | 25 | 27 | 52 | |||
| CA199 (kU/L) | 0.049 | |||||
| ≤37 | 32 | 39 | 71 | 0.352(0.121-1.020) | ||
| >37 | 14 | 6 | 20 | |||
| Tumor size (cm) | 0.040 | |||||
| <5 | 18 | 9 | 27 | 2.621(1.029-6.674) | ||
| ≥5 | 29 | 38 | 67 | |||
| Portal vein tumor thrombus | 0.765 | |||||
| No | 40 | 41 | 81 | 0.836(0.258-2.706) | ||
| Yes | 7 | 6 | 13 | |||
| Embolus | 0.280 | |||||
| No | 33 | 28 | 61 | 1.599(0.681-3.759) | ||
| Yes | 14 | 19 | 33 | |||
| Metastasis | 0.049 | |||||
| No | 43 | 36 | 79 | 3.285(0.963-11.205) | ||
| Yes | 4 | 11 | 15 | |||
| TNM stage | 0.672 | |||||
| Ⅰ/Ⅱ | 28 | 26 | 54 | 1.197(0.521-2.747) | ||
| Ⅲ/Ⅳ | 18 | 20 | 38 | |||
| Recurrence | 0.536 | |||||
| No | 23 | 26 | 49 | 0.774(0.344-1.742) | ||
| Yes | 24 | 21 | 45 | |||
Cell line and cell culture
The human HCC cell line SMMC-7721 used in this study was preserved in the Central Laboratory of Guangxi Medical University (Nanning, Guangxi Zhuang Autonomous Region). The culture medium for SMMC-7721 cells was composed of Eagle medium modified by Dulbecco (DMEM, Thermo Fisher Scientific, USA), 10% fetal bovine serum (FBS, Gibco, Grand Island, NY, USA) and 1% antibiotics (penicillin, 100 µ/ml; streptomycin sulfates,100 mg/ml). The cells were cultured in a constant temperature incubator containing 37oC, 5% CO2.
Cell transfection
SMMC-7721 cells were seeded into 6-well plates at approximately 2×105 cells per well, miR-381-3p mimic, mimic negative control (NC), miR-381-3p inhibitor, and inhibitor negative control were via Tiangen Biochemical Technology Co synthesis. According to the manufacturer's instructions, they were mixed with Lipofectamine 6000 (Invitrogen, Thermo Fisher Scientific, Inc., USA) and Opti-MEM (Gibco, Thermo Fisher Scientific Inc., USA) for SMMC- 7721 cells were transfected.
RNA extraction, reverse transcription and quantitative real-time PCR
Total RNA in 94 tissues and transfected cells was extracted using TRIzol (Invitrogen, Carlsbad, CA, USA), and the total RNA was reverse transcribed into cDNA by miRcute miRNA cDNA kit (Tiangen Biotech, Beijing, Co, Ltd.), miRcute miRNA qPCR kit (Tiangen Biotech, Beijing, Co, Ltd.) was subjected to qRT-PCR.
MTT proliferation assay
The proliferation of HCC cells was detected by Thiazolyl Blue Tetrazolium Bromide (MTT). 1.5×103 transfected HCC cells were seeded into 96-well plates. At the time of culture for 24 h, 48 h, 72 h and 96 h, the original medium was removed, and the mixture of 100µL cell medium and 20µL MTT solution (Sigma-Aldrich, St Louis, MO, USA) was added to each well. After 4 h incubation, the mixture was removed and 100µL of dimethyl sulfoxide (DMSO, Sigma-Aldrich; Shanghai, China) was added and incubating in dark for 15 min, the optical density (OD) value of each well at a wavelength of 490 nm was measured to estimate the number of SMMC-7721 cells.
Transwell assay
The invasive ability of HCC cells was determined by transwell assay. Matrigel and serum-free DMED were mixed in a ratio of 1:10, 60µL mixture was placed in the upper chamber, about 5×104 transfected cells were added after coagulation, and 700µL medium containing 10% serum was added to the lower chamber. After incubation for 48 h, the excess cells on the surface of the membrane were removed. The paraformaldehyde was fixed for 1 h, stained with 0.1% crystal violet for 30 min, washed with PBS, and counted the cells under a microscope.
Wound healing assay
The migration ability of SMMC-7721 cells was evaluated by wound healing assay. 2×105 cells were seeded into 6-well plates, after 48 h transfection, using 200µL tip to scrape transparent straight lines vertically through the cells, the medium was aspirated and washed with phosphate buffered saline (PBS, Gibco). The cells were continuously cultured in a medium without serum. The images were collected at 0 h, 12 h, 24 h and 48 h after the rinsing to observe the cell migration.
Prediction and functional analysis of target genes
Using bioinformatics technology to predict the target genes of miR-381-3p, and 4 databases were selected for prediction, including starBase v2.0 (http://starbase.sysu.edu.cn/starbase2/index.php). Target Scan Human 7.2 (http://www.targetscan.org/vert_72/), miRDB (http://mirdb.org/), DIANA TOOLS - microT-CDS (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=microT_C DS/index). In order to improve the prediction accuracy selected target genes can be predicted by three or more databases. The target gene was subjected to GO (Gene Ontology) enrichment analysis and KEGG (Kyoto Encyclopedia of Genes and Genomes) biochannel enrichment analysis using the DAVID online analysis tool (https://david.ncifcrf.gov/).
Statistical analysis
SPSS 22.0 was used for statistical analysis, and GraphPad Prism 7.0 was used for image mapping. The relationship between miR-381-3p and clinical pathological features of HCC was evaluated by chi-square test, and two independent groups were compared by Student's t-test in the study. P <0.05 was considered statistically significant.
ResultsTop
Low expression of miR-381-3p in HCC
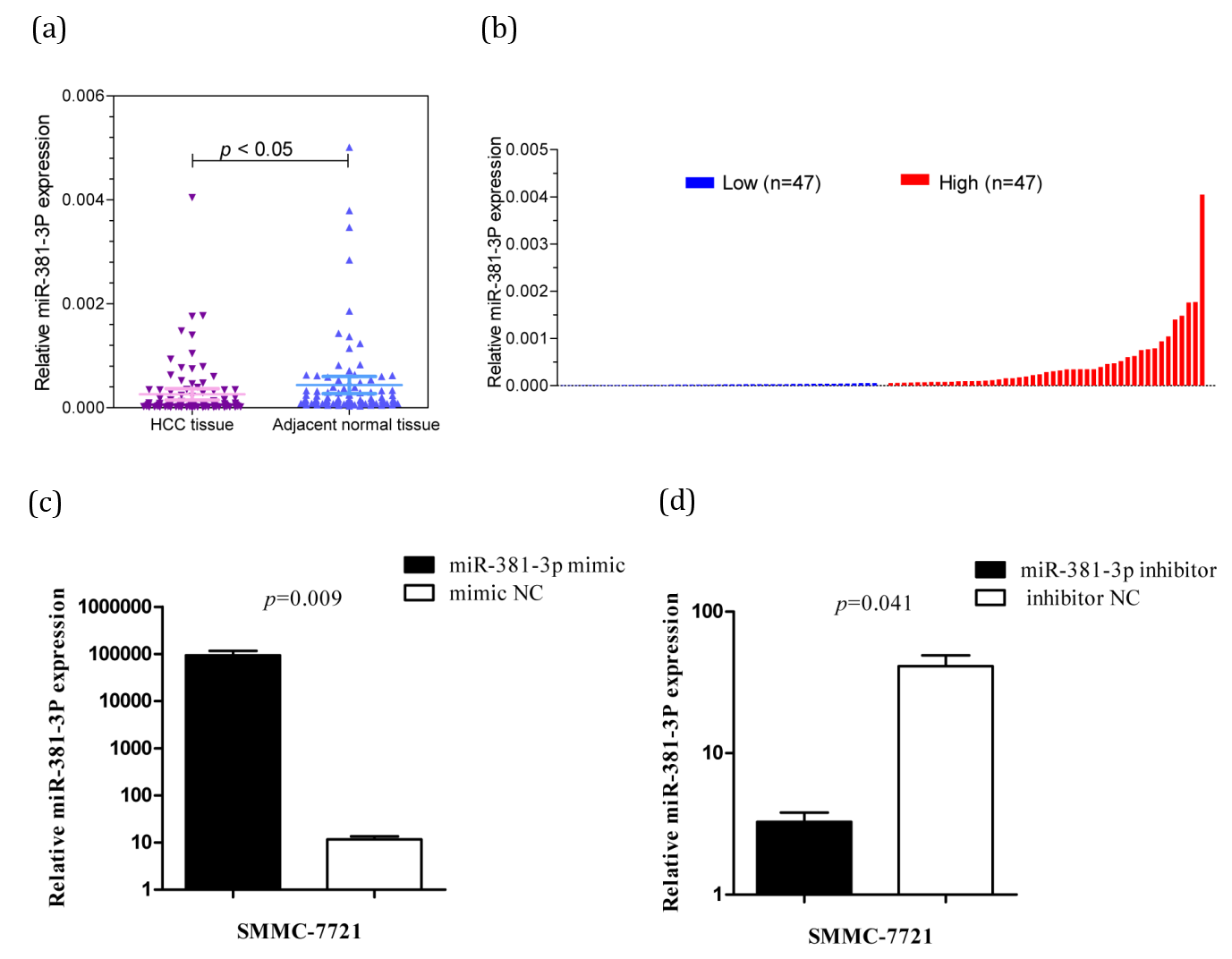
The results showed that the expression of miR-381-3p in HCC tissues was different from that in paracancerous tissues, and it was obviously that miR-381-3p was lowly expressed in HCC tissues (p=0.038) (Figure 1a).
The correlation between the expression of miR-381-3p and the clinicopathological characteristics of HCC patients
We ranked the expression of miR-381-3p in the tissues of 94 HCC patients and grouped them by the median, 94 HCC patients were divided into the high expression group and the low expression group of miR-381-3p (Figure 1b). Statistical analysis showed that the expression of miR-381-3p was associated with gender (p=0.036), CA199 (p=0.049), tumor size (p=0.040), metastasis (p=0.049), but not with age, HBV-DNA, AFP, Portal vein tumor thrombus, embolus, TNM stage, recurrence and other clinicopathological features (Table 1).
Transfection efficiency of SMMC-7721 HCC cells
The expression of miR-381-3p in SMMC-7721 HCC cells transfected with miR-381-3p mimics was higher than that in the mimic NC group (p<0.05) (Figure 1c). The miR-381-3p expression of inhibitor group was lower than that of the inhibitor NC group (p<0.05) (Figure 1d).
miR-381-3p inhibits proliferation of SMMC-7721 HCC cells
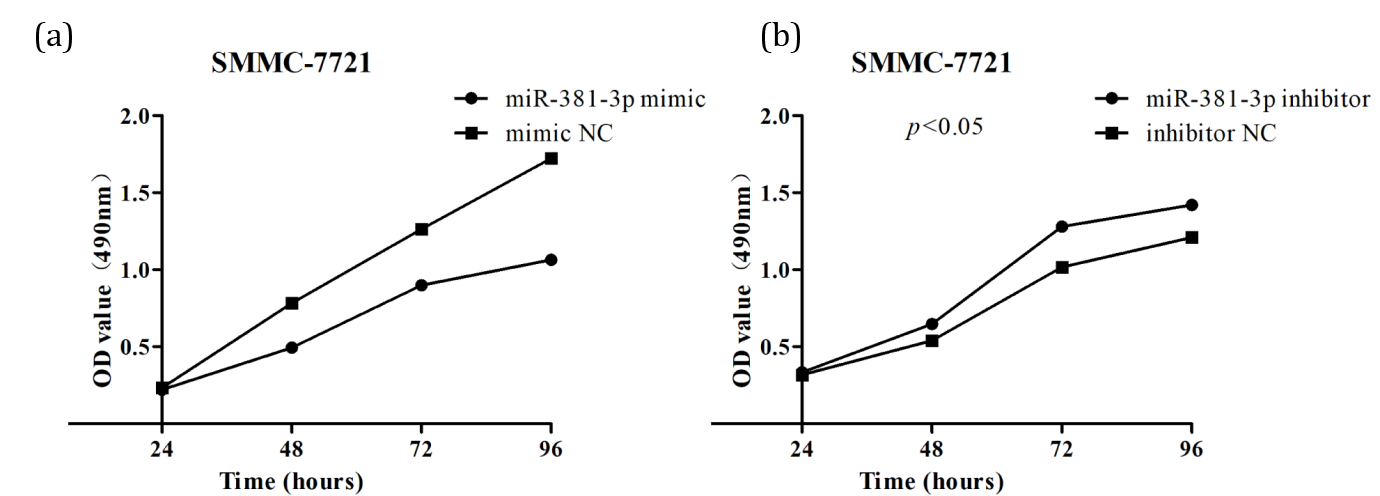
The OD values measured at 48 h, 72 h, and 96 h showed that the proliferation of mimic group cells was significantly lower than that of mimic NC cells (p<0.05) (Figure 2a). At the same time, the proliferation of the inhibitor group cells was significantly higher than that of the inhibitor NC group cells (p<0.05) (Figure 2b). The results showed that miR-381-3p affects the HCC cells proliferation and plays an inhibitory role.
miR-381-3p inhibits invasion and migration of SMMC-7721 HCC cells
The results of transwell assay were shown in Figure 3a. At 48 h of invasion, the number of invasive cells in the mimic group was than that in the mimic NC group (p<0.05) (Figure 3b). Meanwhile, compared with inhibitor NC group, the number of invasive cells in inhibitor group was increased (p<0.05) (Figure 3c).
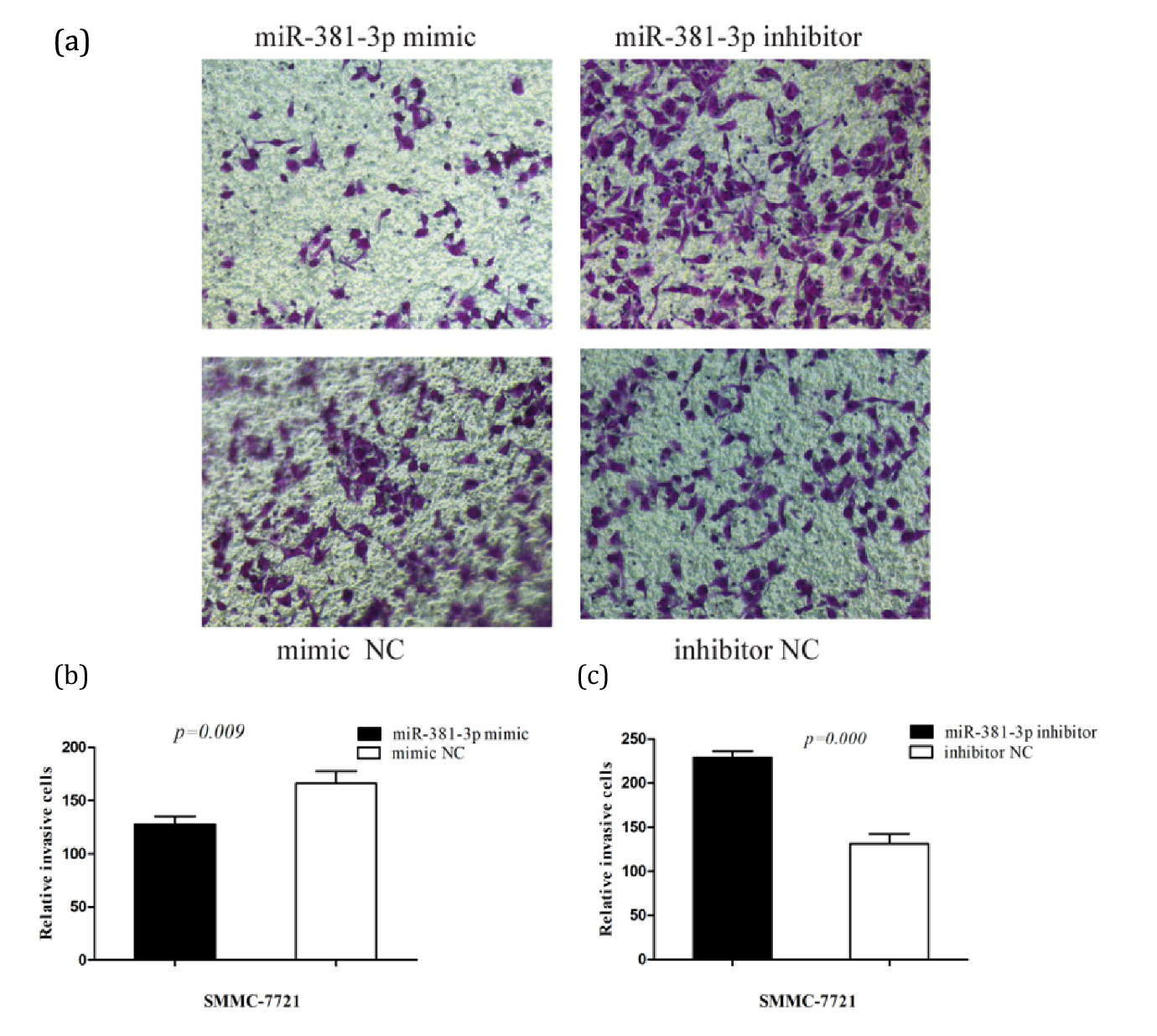
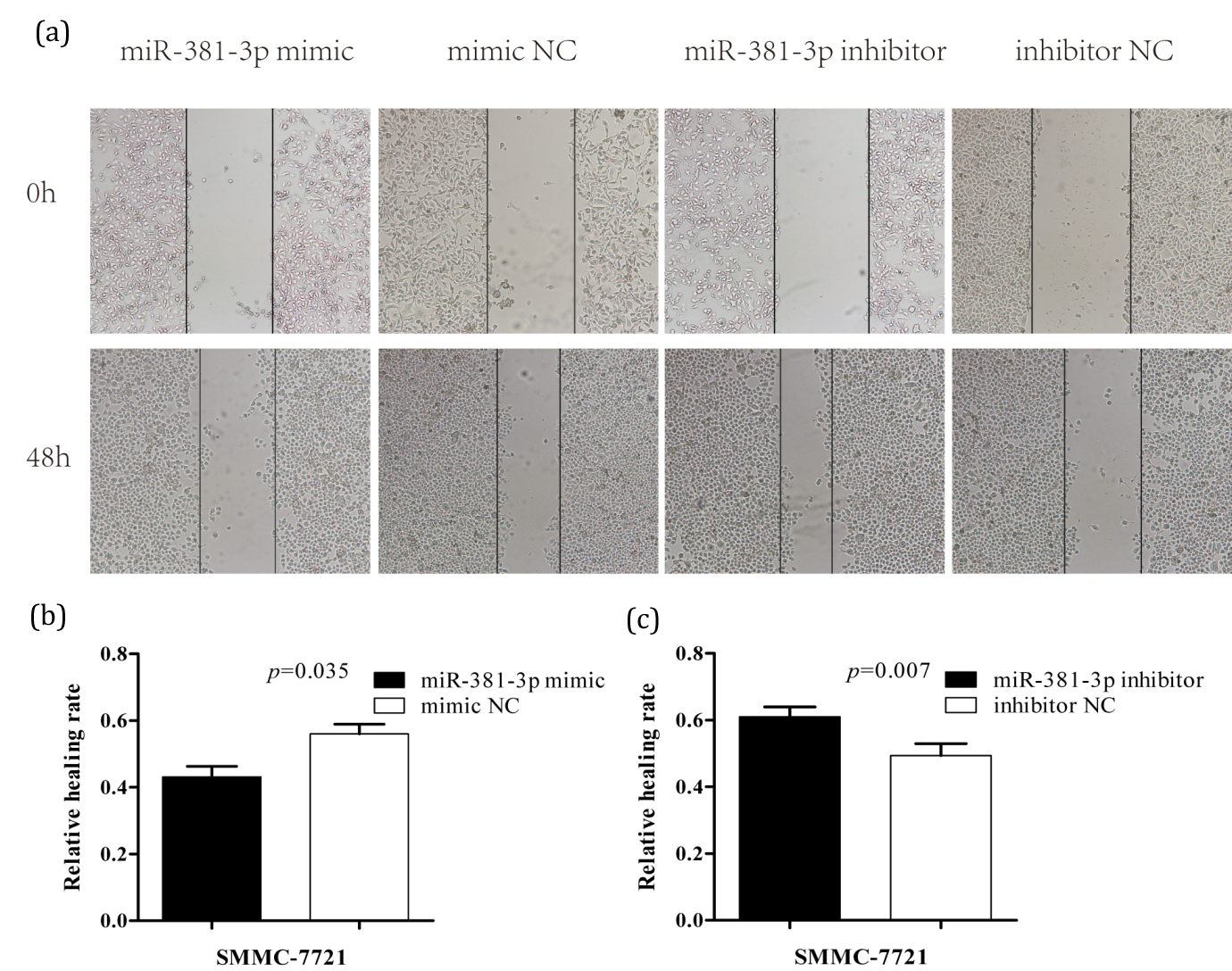
Transwell results demonstrated that miR-381-3p inhibits invasion of HCC cells. The results of wound healing assay were shown in (Figure 4a). At 48 h, the scratch distance in mimic group was larger than that in NC group, and the migration rate and healing rate were smaller (p<0.05) (Figure 4b). However, the scratch distance of the inhibitor group was smaller and the migration and healing rate was greater than that of the inhibitor NC group (p<0.05) (Figure 4c). The wound healing assay results indicated that miR-381-3p inhibited the migration of HCC cells.
Prediction and functional analysis of target genes of miR-381-3p
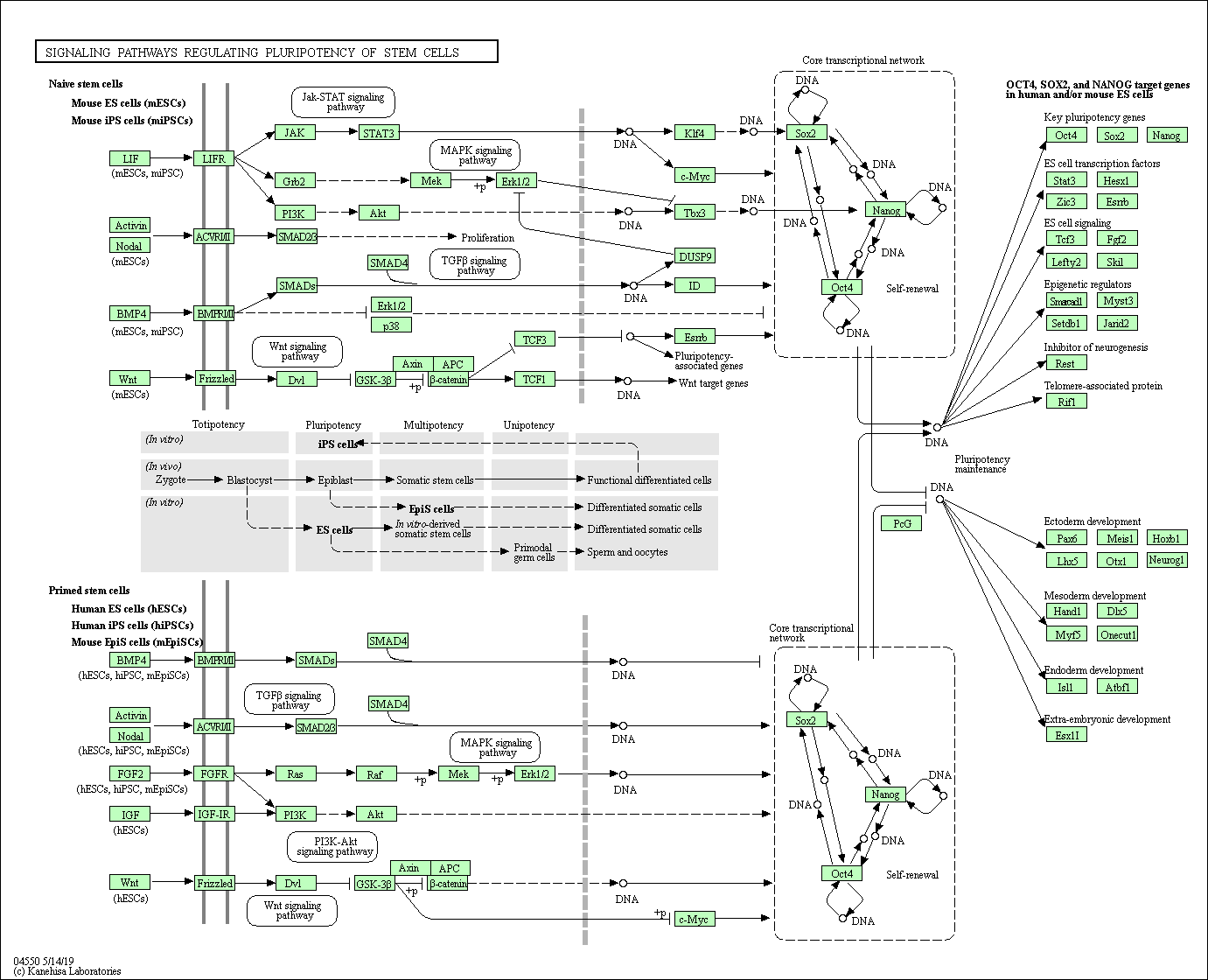
Using the databases, we predicted 854 possible target genes of miR-381-3p in this study. GO enrichment analysis was performed on the predicted target genes of miR-381-3p using the DAVID online analysis tool. The enrichment of the target genes in molecular function (MF), cellular component (CC), and biological process (BP) were ranked according to the p value from low to high, and the top 10 results are shown in Figure 5 (p<0.05). The KEGG pathway enrichment analysis of these genes showed that the target genes of miR-381-3p were enriched in 17 pathways (p<0.01). These pathways were ranked according to p value, and the top 10 results were shown in Figure 6 (p<0.05). The strongest enrichment pathway is signaling pathways regulating pluripotency of stem cells (Figure 7).


DiscussionTop
HCC is the second most common malignant tumor in the world, second only to pancreatic cancer [13]. Viral infections, alcoholic liver disease and nonalcoholic steatohepatitis are representative causes of HCC [14]. It is crucial to find new ways to reduce the mortality rate of HCC and improve the prognosis of patients with HCC. MicroRNAs (miRNAs) can bind to their corresponding target mRNAs, inhibit the function of mRNA, and promote the degradation of mRNA. Through this post-transcriptional regulation mechanism, gene expression can be changed, which has been proved of great significance in a variety of organisms [15, 16]. Abnormal miRNAs expression can affect a variety of human life activities, and is associated with human diseases, including cancer [17]. These changes in miRNA often affect cell proliferation and apoptosis, which affect the occurrence and development of cancer. Some miRNAs are oncogenes, some are anti-oncogene, and play an important role in different steps of tumorigenesis, from initiation to development and even metastasis [18, 19]. Different miRNAs can affect the same cancer, and one miRNA can also play a role in different cancers [20]. Studies have shown that dysregulation of miRNAs interferes with the normal cell cycle, which can cause HCC [21]. Therefore, by studying the biological functions of miRNAs affecting HCC and discovering the regulation of miRNAs in HCC, miRNAs may become a new diagnostic and therapeutic tool, which is especially important for patients with HCC [22, 23].
In the past, there have been many studies on the role of miR-381-3p in different cancers. Shang et al. [24] found that miR-381-3p expression was down-regulated in cervical cancer tissues and inhibited cell proliferation, migration and invasion, its expression also promoted cell cycle arrest and accelerates apoptosis. Li et al. [25] studied the effect of miR-381-3p on bladder cancer and found that the low miR-381-3p expression was in bladder cancer tissues, and the proliferation and migration of bladder cancer cells were inhibited. Kong [26] demonstrated that miR-381-3p expression was low in papillary thyroid carcinoma tissues and cell lines, and miR-381-3p inhibited PTC Cell proliferation, migration and invasion.
In our study, the expression of miR-381-3p was significantly down-regulated in HCC tissues, and it was correlated with the clinical and pathological characteristics of HCC patients, including gender, CA199, tumor size and metastasis. Cell function assays showed that miR-381-3p can inhibit the proliferation, migration and invasion of SMMC-7721 HCC cells. In contrast to previous studies on miR-381-3p [14-17], it can be found that miR-381-3p inhibited multiple cancers and mainly affected the proliferation and migration of cancer cells. All the results suggest that miR-381-3p may be a potential target for clinical treatment of HCC. Although the study has predicted the target genes of miR-381-3p by bioinformatics methods and analyzed their functions and pathways, biological assays are still needed to determine the target genes and their binding sites regulated by miR-381-3p. Therefore, the mechanism of miR-381-3p in HCC can be further explored in future studies.
ConclusionTop
Our results indicate that miR-381 is low expressed in HCC and inhibits proliferation, invasion and migration of HCC cells. MiR-381-3p may be a new target for the future treatment of HCC.
Conflicts of interest
Authors declare no conflict of interest.
ReferencesTop
[1]Kulik L, El-Serag HB. Epidemiology and management of hepatocellular carcinoma. Gastroenterology. 2019; 156(2):477−491.Article Pubmed
[2]Zhong JH, Peng NF, You XM, Ma L, Xiang X, et al. Tumor stage and primary treatment of hepatocellular carcinoma at a large tertiary hospital in China: A real-world study. Oncotarget. 2017; 8(11):18296−18302.Article Pubmed
[3]Ozakyol A. Global epidemiology of hepatocellular carcinoma (HCC Epidemiology). J Gastrointest Cancer. 2017; 48(3):238−240.Article Pubmed
[4]Zhu ZX, Huang JW, Liao MH, Zeng Y. Treatment strategy for hepatocellular carcinoma in China: radiofrequency ablation versus liver resection. Jpn J Clin Oncol. 2016; 46(12):1075−1080.Article Pubmed
[5]Zhi Y, Abudoureyimu M, Zhou H, Wang T, Feng B, et al. FOXM1-Mediated LINC-ROR regulates the proliferation and sensitivity to sorafenib in hepatocellular carcinoma. Mol Ther Nucleic Acids. 2019; 16:576−588.Article Pubmed
[6]Simonson B, Das S. MicroRNA therapeutics: the next magic bullet?. Mini Rev Med Chem. 2015; 15(6):467−474.Article Pubmed
[7]Acunzo M, Romano G, Wernicke D, Croce CM. MicroRNA and cancer--a brief overview. Adv Biol Regul. 2015; 57:1−9.Article Pubmed
[8]Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014; 20(8):460−469.Article Pubmed
[9]Ruan L, Chen J, Ruan L, Yang T, Wang P. MicroRNA-186 suppresses lung cancer progression by targeting SIRT6. Cancer Biomark. 2018; 21(2):415−423.Article Pubmed
[10]Wu D, Niu X, Pan H, Zhang Z, Zhou Y, et al. MicroRNA-497 targets hepatoma-derived growth factor and suppresses human prostate cancer cell motility. Mol Med Rep. 2016; 13(3):2287−2292.Article Pubmed
[11]Hayes CN, Chayama K. MicroRNAs as biomarkers for liver disease and hepatocellular carcinoma. Int J Mol Sci. 2016; 17(3):280.Article Pubmed
[12]Wong QW, Lung RW, Law PT, Lai PB, Chan KY, et al. MicroRNA-223 is commonly repressed in hepatocellular carcinoma and potentiates expression of Stathmin1. Gastroenterology. 2008; 135(1):257−269.Article Pubmed
[13]Villanueva A. Hepatocellular carcinoma. N Engl J Med. 2019; 380(15):1450−1462.Article Pubmed
[14]Tsuchiya A, Ogawa M, Watanabe T, Takeuchi S, Kojima Y, et al. Diverse perspectives to address for the future treatment of heterogeneous hepatocellular carcinoma. Heliyon. 2019; 5(3):e01325.Article Pubmed
[15]Wu KL, Tsai YM, Lien CT, Kuo PL, Huang AJ. The roles of microRNA in lung cancer. Int J Mol Sci. 2019; 20(7):1611.Article Pubmed
[16]Remsburg C, Konrad K, Sampilo NF, Song JL. Analysis of microRNA functions. Methods Cell Biol. 2019; 151:323−334.Article Pubmed
[17]Costa PM, Pedroso De Lima MC. MicroRNAs as molecular targets for cancer therapy: on the modulation of microRNA expression. Pharmaceuticals (Basel). 2013; 6(10):1195−1220.Article Pubmed
[18]Neelakandan K, Babu P, Nair S. Emerging roles for modulation of microRNA signatures in cancer chemoprevention. Curr Cancer Drug Targets. 2012; 12(6):716−740.Article Pubmed
[19]Xi JJ. MicroRNAs in cancer. Cancer Treat Res. 2013; 158:119−137.Article Pubmed
[20]Zhang B, Pan X, Cobb GP, Anderson TA. MicroRNAs as oncogenes and tumor suppressors. Dev Biol. 2007; 302(1):1−12.Article Pubmed
[21]Chen E, Xu X, Liu R, Liu T. Small but heavy role: Micrornas in hepatocellular carcinoma progression. 2018; 2018:6784607.Article Pubmed
[22]Berger F, Reiser MF. Micro-RNAs as potential new molecular biomarkers in oncology: have they reached relevance for the clinical imaging sciences. Theranostics. 2013; 3(12):943−952.Article Pubmed
[23]Chaiteerakij R, Addissie BD, Roberts LR. Update on biomarkers of hepatocellular carcinoma. Clin Gastroenterol Hepatol. 2015; 13(2):237−245.Article Pubmed
[24]Shang A, Zhou C, Bian G, Chen W, Lu W, et al. miR-381-3p restrains cervical cancer progression by downregulating FGF7. 2019; 120(1):778−789.Article Pubmed
[25]Li J, Ying Y, Xie H, Jin K, Yan H, et al. Dual regulatory role of CCNA2 in modulating CDK6 and MET-mediated cell-cycle pathway and EMT progression is blocked by miR-381-3p in bladder cancer. FASEB J. 2019; 33(1):1374−1388.Article Pubmed
[26]Kong W, Yang L, Li PP, Kong QQ, Wang HY, et al. MiR-381-3p inhibits proliferation, migration and invasion by targeting LRP6 in papillary thyroid carcinoma. Eur Rev Med Pharmacol Sci. 2018; 22(12):3804−3811.Article Pubmed


